
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,824,650 – 14,825,007 |
| Length | 357 |
| Max. P | 0.979265 |

| Location | 14,824,650 – 14,824,770 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -29.40 |
| Consensus MFE | -27.69 |
| Energy contribution | -27.80 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.657323 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14824650 120 + 27905053 AGUUUUCCUAUGCAAAUCUGAGCCGCAUUCGCAUUCCCGACAAAUUUUUGGGAAAGUUGACACAAUUUGAACCCCUUCAACCAGCUUUGGCCAGUCAGGCAUUCAGAUGCUCUGAGACAA .(((((.....(((..((((((((((....)).(((((((.......)))))))...((((.....(((((....)))))((......))...)))))))..))))))))...))))).. ( -30.00) >DroSec_CAF1 2818 120 + 1 AGUUUUCCUAUGCAAAUCUGAGCCGCAUUCGCAUUCCCGACAAAUUUUUGGGAAAGUUGACACAAUUUGAACCCCUUCAACCAGCUUUGGCCAGUCAGCCAUUCAGAUGCUCUGAAACAA .(((((.....(((..((((((..((....)).(((((((.......))))))).((((((.....(((((....)))))((......))...))))))..)))))))))...))))).. ( -29.10) >DroSim_CAF1 2952 120 + 1 AGUUUUCCUAUGCAAAUCUGAGCCGCAUUCGCAUUCCCGACAAAUUUUUGGGAAAGUUGACACAAUUUGAACCCCUUCAACCAGCUUUGGCCAGUCAGCCAUUCAGAUGCUCUGAAACAA .(((((.....(((..((((((..((....)).(((((((.......))))))).((((((.....(((((....)))))((......))...))))))..)))))))))...))))).. ( -29.10) >consensus AGUUUUCCUAUGCAAAUCUGAGCCGCAUUCGCAUUCCCGACAAAUUUUUGGGAAAGUUGACACAAUUUGAACCCCUUCAACCAGCUUUGGCCAGUCAGCCAUUCAGAUGCUCUGAAACAA .(((((.....(((..((((((..((....)).(((((((.......))))))).((((((.....(((((....)))))((......))...))))))..)))))))))...))))).. (-27.69 = -27.80 + 0.11)



| Location | 14,824,690 – 14,824,810 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.28 |
| Mean single sequence MFE | -39.50 |
| Consensus MFE | -37.89 |
| Energy contribution | -37.57 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979265 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14824690 120 - 27905053 UGCUGCUCACGGUUUUAGCCCCAAAGACCGGCGAUGUUAUUUGUCUCAGAGCAUCUGAAUGCCUGACUGGCCAAAGCUGGUUGAAGGGGUUCAAAUUGUGUCAACUUUCCCAAAAAUUUG ...((..((((((((.((((((...(((((((((((((...........)))))).....(((.....)))....)))))))...)))))).)))))))).))................. ( -39.40) >DroSec_CAF1 2858 117 - 1 ---UGCUCACGGUUUUAGCCCCAAAGACCAGUGAUGUUAUUUGUUUCAGAGCAUCUGAAUGGCUGACUGGCCAAAGCUGGUUGAAGGGGUUCAAAUUGUGUCAACUUUCCCAAAAAUUUG ---((..((((((((.((((((...(((((((............(((((.....)))))(((((....)))))..)))))))...)))))).)))))))).))................. ( -40.50) >DroSim_CAF1 2992 120 - 1 CGCUGCUCACGGUUUUAGUCCCAAAGACCAGCGAUGUUAUUUGUUUCAGAGCAUCUGAAUGGCUGACUGGCCAAAGCUGGUUGAAGGGGUUCAAAUUGUGUCAACUUUCCCAAAAAUUUG ...((..((((((((.(..(((...(((((((............(((((.....)))))(((((....)))))..)))))))...)))..).)))))))).))................. ( -38.60) >consensus _GCUGCUCACGGUUUUAGCCCCAAAGACCAGCGAUGUUAUUUGUUUCAGAGCAUCUGAAUGGCUGACUGGCCAAAGCUGGUUGAAGGGGUUCAAAUUGUGUCAACUUUCCCAAAAAUUUG ...((..((((((((.((((((...(((((((((((((...........))))))....(((((....)))))..)))))))...)))))).)))))))).))................. (-37.89 = -37.57 + -0.33)


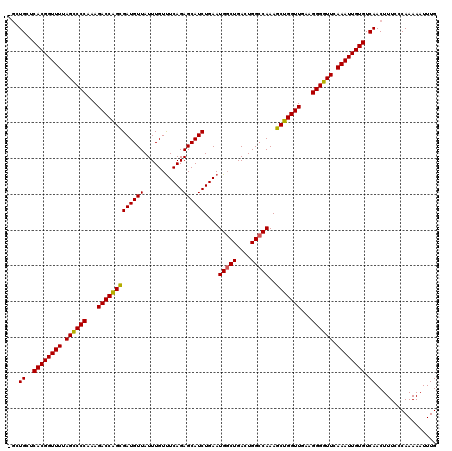
| Location | 14,824,810 – 14,824,927 |
|---|---|
| Length | 117 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.92 |
| Mean single sequence MFE | -41.80 |
| Consensus MFE | -41.77 |
| Energy contribution | -41.77 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.937679 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14824810 117 + 27905053 U---UGUGCCUUUCACUUGGCCAGGCUCUUAGCCCGGAGUCCCAAAUGGCCAACACCUUGCCAACACCAGACACACUCGGCUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCA .---((.(((...((..(((((((((..((((((.((.(((.....((((.........))))......)))...)).)))))).....)))))))))..))..((((...))))))))) ( -41.80) >DroSec_CAF1 2975 115 + 1 -----GUGCCUUUCACUUGGCCAGGCUCUUAGCCCGGAGUCCCAAAUGGCCAACACCUUGCCAACACCAGACACACUCGGCUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCA -----(.(((...((..(((((((((..((((((.((.(((.....((((.........))))......)))...)).)))))).....)))))))))..))..((((...)))))))). ( -41.70) >DroSim_CAF1 3112 120 + 1 UAGUAGUGCCUUUCACUUGGCCAGGCUCUUAGCCCGGAGUCCCAAAUGGCCAACACCUUGCCAACACCAGACACACUCGGCUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCA .....(.(((...((..(((((((((..((((((.((.(((.....((((.........))))......)))...)).)))))).....)))))))))..))..((((...)))))))). ( -41.90) >consensus U____GUGCCUUUCACUUGGCCAGGCUCUUAGCCCGGAGUCCCAAAUGGCCAACACCUUGCCAACACCAGACACACUCGGCUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCA .....(.(((...((..(((((((((..((((((.((.(((.....((((.........))))......)))...)).)))))).....)))))))))..))..((((...)))))))). (-41.77 = -41.77 + 0.00)



| Location | 14,824,810 – 14,824,927 |
|---|---|
| Length | 117 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.92 |
| Mean single sequence MFE | -50.57 |
| Consensus MFE | -50.20 |
| Energy contribution | -50.20 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.834312 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14824810 117 - 27905053 UGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAGCCGAGUGUGUCUGGUGUUGGCAAGGUGUUGGCCAUUUGGGACUCCGGGCUAAGAGCCUGGCCAAGUGAAAGGCACA---A (((((..(((....(((.(((((((((((.((((.....))))....((((.......)))).....)))))))))))))).)))((((.....))))))))).(((.....))).---. ( -50.40) >DroSec_CAF1 2975 115 - 1 UGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAGCCGAGUGUGUCUGGUGUUGGCAAGGUGUUGGCCAUUUGGGACUCCGGGCUAAGAGCCUGGCCAAGUGAAAGGCAC----- ..(((((((...))))(((...(((((((((.....((((((..(.(.((((.(...((((.........))))..).)))).)).))))))..))))))))).)))...)))..----- ( -50.20) >DroSim_CAF1 3112 120 - 1 UGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAGCCGAGUGUGUCUGGUGUUGGCAAGGUGUUGGCCAUUUGGGACUCCGGGCUAAGAGCCUGGCCAAGUGAAAGGCACUACUA (((((..(((....(((.(((((((((((.((((.....))))....((((.......)))).....)))))))))))))).)))((((.....)))))))))((((.....)))).... ( -51.10) >consensus UGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAGCCGAGUGUGUCUGGUGUUGGCAAGGUGUUGGCCAUUUGGGACUCCGGGCUAAGAGCCUGGCCAAGUGAAAGGCAC____A ..(((((((...))))(((...(((((((((.....((((((..(.(.((((.(...((((.........))))..).)))).)).))))))..))))))))).)))...)))....... (-50.20 = -50.20 + -0.00)



| Location | 14,824,887 – 14,825,007 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -36.20 |
| Consensus MFE | -34.59 |
| Energy contribution | -34.70 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.762616 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14824887 120 + 27905053 CUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCACAAUCUUGACAUUAUUCUCCAUGGAGAGCUCUUCGCGCGGCGACAAUGCGGAAAAACAACUUUCGCAAAUUCUUUCAAUU ....(((((((.(((((.......((((...)))))))))...............((((....))))((.....)))))))))...(((((((.......)))))))............. ( -36.01) >DroSec_CAF1 3050 120 + 1 CUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCACAAUUUUGACAUUAUUCUCCAUGGAGAGCUCCUCGCGCGGCGACAAUGCGGAAAAACAACUUUCGCAAAUUCUUUCAAUU ....(((((((...((.....)).((((((..((((....)........(((........))))))..))))))..)))))))...(((((((.......)))))))............. ( -37.10) >DroSim_CAF1 3192 120 + 1 CUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCACAAUCUUGACAUUAUUCUCCAUGGUGAGCUCCUCGCGCGGCGACAAUGCGGAAAAACAACUUUCGCAAAUUCUUUCAAUU ....(((((((...((.....)).((((((..((((.(((.......((......))....)))))))))))))..)))))))...(((((((.......)))))))............. ( -35.50) >consensus CUAAUCGCCGCCUGGCCAUCUGCGGAGGAGUCCUCGGCCACAAUCUUGACAUUAUUCUCCAUGGAGAGCUCCUCGCGCGGCGACAAUGCGGAAAAACAACUUUCGCAAAUUCUUUCAAUU ....(((((((...((.....)).((((((..((((....)........(((........))))))..))))))..)))))))...(((((((.......)))))))............. (-34.59 = -34.70 + 0.11)



| Location | 14,824,887 – 14,825,007 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -40.80 |
| Consensus MFE | -39.37 |
| Energy contribution | -40.03 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.539679 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14824887 120 - 27905053 AAUUGAAAGAAUUUGCGAAAGUUGUUUUUCCGCAUUGUCGCCGCGCGAAGAGCUCUCCAUGGAGAAUAAUGUCAAGAUUGUGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAG ....((((((....((....))..))))))......((((((((((.....))((((....))))...............((((((.(((....))).....)))))).))))))))... ( -38.40) >DroSec_CAF1 3050 120 - 1 AAUUGAAAGAAUUUGCGAAAGUUGUUUUUCCGCAUUGUCGCCGCGCGAGGAGCUCUCCAUGGAGAAUAAUGUCAAAAUUGUGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAG ....((((((....((....))..))))))...((((((((((((.((((((..(((...((...(((((......)))))..)))))..))))))))).(.....).)))))))))... ( -43.60) >DroSim_CAF1 3192 120 - 1 AAUUGAAAGAAUUUGCGAAAGUUGUUUUUCCGCAUUGUCGCCGCGCGAGGAGCUCACCAUGGAGAAUAAUGUCAAGAUUGUGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAG ....((((((....((....))..))))))...((((((((((((.((((((....((.(((...(((((......)))))..))).)).))))))))).(.....).)))))))))... ( -40.40) >consensus AAUUGAAAGAAUUUGCGAAAGUUGUUUUUCCGCAUUGUCGCCGCGCGAGGAGCUCUCCAUGGAGAAUAAUGUCAAGAUUGUGGCCGAGGACUCCUCCGCAGAUGGCCAGGCGGCGAUUAG ....((((((....((....))..))))))...((((((((((((.((((((...(((.(((...(((((......)))))..))).)))))))))))).(.....).)))))))))... (-39.37 = -40.03 + 0.67)


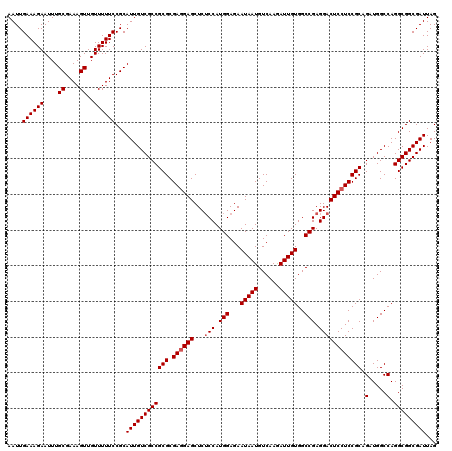
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:59:53 2006