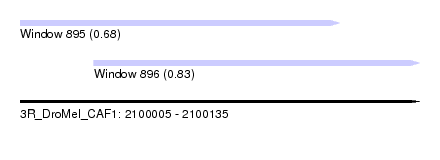

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 2,100,005 – 2,100,135 |
| Length | 130 |
| Max. P | 0.826614 |

| Location | 2,100,005 – 2,100,109 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 86.54 |
| Mean single sequence MFE | -15.77 |
| Consensus MFE | -11.70 |
| Energy contribution | -12.04 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.681958 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
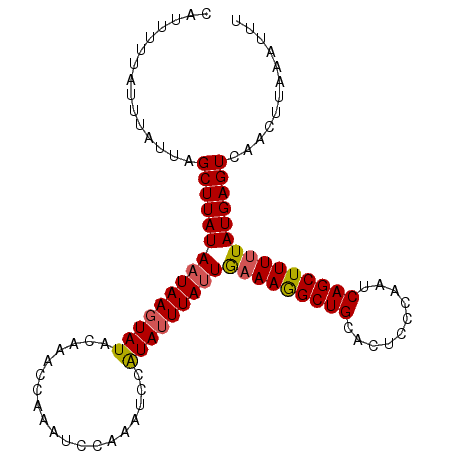
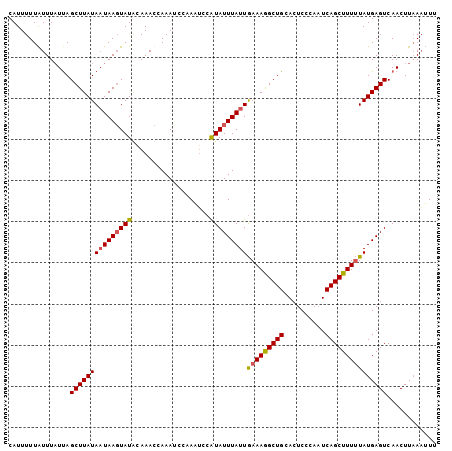
>3R_DroMel_CAF1 2100005 104 + 27905053 CAUUUUUAUUUAUUAGCUUAUAAUAACUAUACAAAUCAAAGCCUAAGCCAUAUUUAUUGAAAAGCUGCACUCCCAUUCAGCUUUUUAUGAGUCAACUUAACUUU .........((((((.....))))))...........((((..((((....((((((.(((((((((..........)))))))))))))))...)))).)))) ( -13.60) >DroSec_CAF1 243000 103 + 1 CAUUUUUGUUUUUUAGCUUAUAAUAAGUAUGGAAACCAAAUCCAAUUCCAUAUUUAUUAAAAGGCUGCACUCCCAAUCAGCUUUAUAUGAGUCAACUUUAAAU- ...............(((((((((((((((((((...........)))))))))))))).(((((((..........)))))))...)))))...........- ( -21.20) >DroSim_CAF1 246084 103 + 1 CAUUUUUAUUUAUCAGCUUAUA-UAAGUAUACAAACCAAAUCCAAAUCCGUAUUUAUUGAAAGGCUGCACUCCCAAUCAGCUUUUUAUGAGUCAACUUAAAUUU .......(((((...(((((((-(((((((...................)))))))).(((((((((..........))))))))))))))).....))))).. ( -12.51) >consensus CAUUUUUAUUUAUUAGCUUAUAAUAAGUAUACAAACCAAAUCCAAAUCCAUAUUUAUUGAAAGGCUGCACUCCCAAUCAGCUUUUUAUGAGUCAACUUAAAUUU ...............(((((((((((((((...................)))))))))(((((((((..........)))))))))))))))............ (-11.70 = -12.04 + 0.34)

| Location | 2,100,029 – 2,100,135 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 85.53 |
| Mean single sequence MFE | -17.91 |
| Consensus MFE | -13.05 |
| Energy contribution | -13.50 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.826614 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
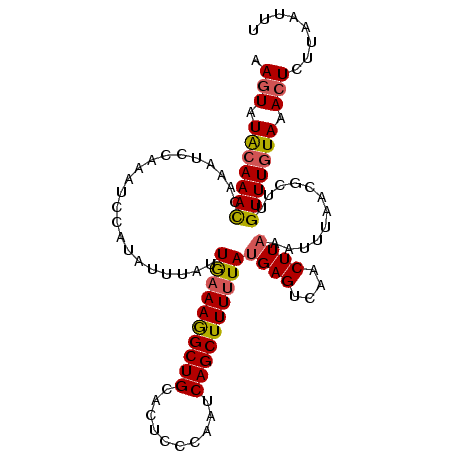
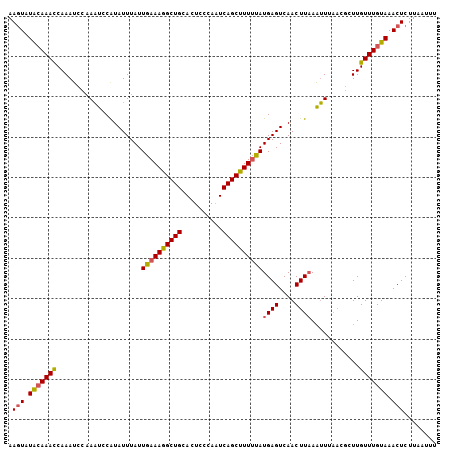
>3R_DroMel_CAF1 2100029 106 + 27905053 AACUAUACAAAUCAAAGCCUAAGCCAUAUUUAUUGAAAAGCUGCACUCCCAUUCAGCUUUUUAUGAGUCAACUUAACUUUAACGCUUGUUUGUAAACUCUUAACUU .....(((((((..((((...............((((((((((..........))))))))))((((....))))........)))))))))))............ ( -18.80) >DroSec_CAF1 243024 100 + 1 AAGUAUGGAAACCAAAUCCAAUUCCAUAUUUAUUAAAAGGCUGCACUCCCAAUCAGCUUUAUAUGAGUCAACUUUAAAU------UUGUUUGUAAACUCUUAAUUU ((((((((((...........)))))))))).....(((((((..........)))))))....((((..((.......------......))..))))....... ( -16.62) >DroSim_CAF1 246107 106 + 1 AAGUAUACAAACCAAAUCCAAAUCCGUAUUUAUUGAAAGGCUGCACUCCCAAUCAGCUUUUUAUGAGUCAACUUAAAUUUAACGCUUGUUUGUAAACUCUUAAUUU .(((.(((((((.....................((((((((((..........))))))))))((((....))))............))))))).)))........ ( -18.30) >consensus AAGUAUACAAACCAAAUCCAAAUCCAUAUUUAUUGAAAGGCUGCACUCCCAAUCAGCUUUUUAUGAGUCAACUUAAAUUUAACGCUUGUUUGUAAACUCUUAAUUU .(((.(((((((.....................((((((((((..........))))))))))((((....))))............))))))).)))........ (-13.05 = -13.50 + 0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:09 2006