
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,762,463 – 14,762,555 |
| Length | 92 |
| Max. P | 0.996255 |

| Location | 14,762,463 – 14,762,555 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 91.74 |
| Mean single sequence MFE | -21.71 |
| Consensus MFE | -17.72 |
| Energy contribution | -18.12 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.67 |
| SVM RNA-class probability | 0.996255 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
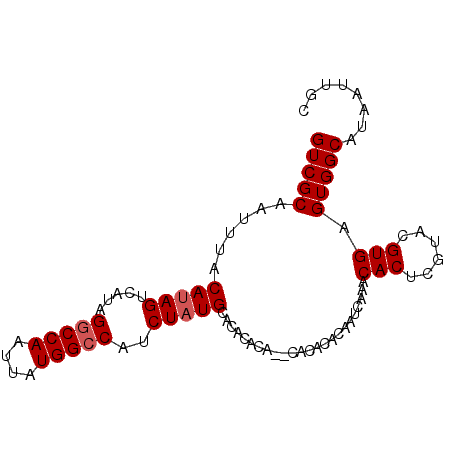
>3R_DroMel_CAF1 14762463 92 + 27905053 GUCGCAAUUUACAUAGUCAUAGGCCAAUUAUGGCCAUCUAUGCACACACACACACACACAAUCAAACACUUGUACGUGAGUGGCAUAAUUGC ...(((((.............(((((....)))))...(((((.(((.(((..(((..............)))..))).))))))))))))) ( -23.04) >DroSec_CAF1 21374 90 + 1 GUCGCAAUUUACAUAGUCAUAGGCCAAUUAUGGCCAUCUAUGCACACACA--CACACACAAUCAAACACUCGUACGUGAGUGGCAUAAUUGC ...(((((.............(((((....)))))...(((((.(((.((--(..((..............))..))).))))))))))))) ( -22.24) >DroSim_CAF1 22534 92 + 1 GUCGCAAUUUACAUAGUCAUAGGCCAAUUAUGGCCAUCUAUGCACACACACACACACACAAUCAAACACUCGUACGUGAGUGGCAUAAUUGC ...(((((.............(((((....)))))...(((((.(((.(((..((................))..))).))))))))))))) ( -22.09) >DroEre_CAF1 21628 83 + 1 GUCGCAAUUUACAUAGUCAUAG-CCAAUUAUGGCCAUCUAUGCACAC--------ACACAAUCAAACACCCGUACGUGAGUGGCAUAAUUGC ...(((((.......(((((((-....)))))))....(((((.(((--------.(((...(........)...))).))))))))))))) ( -18.30) >DroYak_CAF1 22045 84 + 1 GUCGCCAUUUACAUAGUCAUAGGCCAAUUAUGGCCAUCUCUGCACAC--------CCACAAUCAAACACGCGUACGUGAGUGGCAUAAUUGC ...(((((((((.........(((((....)))))......((....--------..............))....)))))))))........ ( -22.87) >consensus GUCGCAAUUUACAUAGUCAUAGGCCAAUUAUGGCCAUCUAUGCACACACA__CACACACAAUCAAACACUCGUACGUGAGUGGCAUAAUUGC (((((......(((((.....(((((....)))))..)))))........................(((......))).)))))........ (-17.72 = -18.12 + 0.40)


| Location | 14,762,463 – 14,762,555 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 91.74 |
| Mean single sequence MFE | -30.96 |
| Consensus MFE | -22.64 |
| Energy contribution | -23.32 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.31 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.956534 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
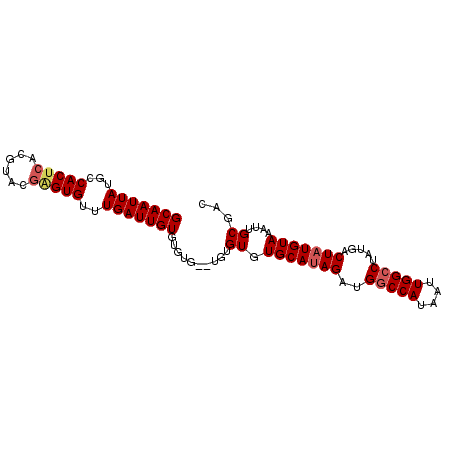
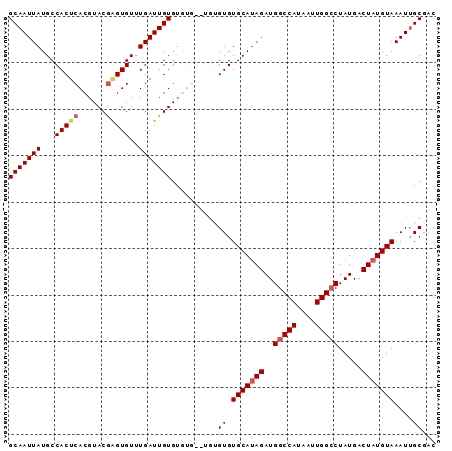
>3R_DroMel_CAF1 14762463 92 - 27905053 GCAAUUAUGCCACUCACGUACAAGUGUUUGAUUGUGUGUGUGUGUGUGUGCAUAGAUGGCCAUAAUUGGCCUAUGACUAUGUAAAUUGCGAC ((((((..(((((.(((((((((........))))))))).))).)).(((((((..(((((....))))).....)))))))))))))... ( -34.30) >DroSec_CAF1 21374 90 - 1 GCAAUUAUGCCACUCACGUACGAGUGUUUGAUUGUGUGUG--UGUGUGUGCAUAGAUGGCCAUAAUUGGCCUAUGACUAUGUAAAUUGCGAC ((((((..(((((.(((((((((........)))))))))--.))).))((((((..(((((....))))).....)))))).))))))... ( -32.90) >DroSim_CAF1 22534 92 - 1 GCAAUUAUGCCACUCACGUACGAGUGUUUGAUUGUGUGUGUGUGUGUGUGCAUAGAUGGCCAUAAUUGGCCUAUGACUAUGUAAAUUGCGAC ((((((..(((((.(((((((((........))))))))).))).)).(((((((..(((((....))))).....)))))))))))))... ( -34.00) >DroEre_CAF1 21628 83 - 1 GCAAUUAUGCCACUCACGUACGGGUGUUUGAUUGUGU--------GUGUGCAUAGAUGGCCAUAAUUGG-CUAUGACUAUGUAAAUUGCGAC (((((((...(((((......)))))..)))))))((--------(..((((((((((((((....)))-))))..)))))))...)))... ( -28.10) >DroYak_CAF1 22045 84 - 1 GCAAUUAUGCCACUCACGUACGCGUGUUUGAUUGUGG--------GUGUGCAGAGAUGGCCAUAAUUGGCCUAUGACUAUGUAAAUGGCGAC .((((((((((((((..((((((.((........)).--------)))))).))).))).))))))))(((....((...))....)))... ( -25.50) >consensus GCAAUUAUGCCACUCACGUACGAGUGUUUGAUUGUGUGUG__UGUGUGUGCAUAGAUGGCCAUAAUUGGCCUAUGACUAUGUAAAUUGCGAC (((((((...(((((......)))))..)))))))..........((.(((((((..(((((....))))).....)))))))....))... (-22.64 = -23.32 + 0.68)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:43 2006