
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,756,412 – 14,756,560 |
| Length | 148 |
| Max. P | 0.981314 |

| Location | 14,756,412 – 14,756,503 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 81.90 |
| Mean single sequence MFE | -28.01 |
| Consensus MFE | -18.94 |
| Energy contribution | -19.80 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.691985 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
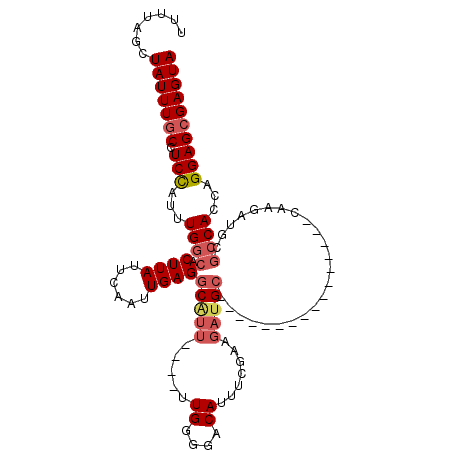
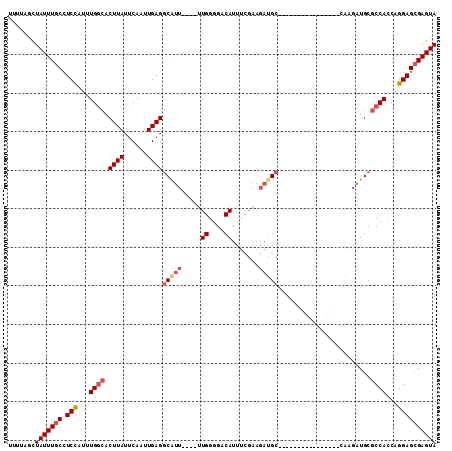
>3R_DroMel_CAF1 14756412 91 - 27905053 UUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUU----UUGGGGACAUUUCGAAGAUGC----------------CAAGAUGCGCCACCAGGAGCGAGUA .......(((((((.(((...((((.((((......))))(((((----((((...((((.....)))))----------------))))))))))))...)))))))))) ( -31.20) >DroSec_CAF1 15371 91 - 1 UUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUU----UUGGGGACAUUUCGAAGAUGC----------------CAAGAUGCGCCACCAGGAGCGAGUA .......(((((((.(((...((((.((((......))))(((((----((((...((((.....)))))----------------))))))))))))...)))))))))) ( -31.20) >DroSim_CAF1 16532 91 - 1 UUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUU----UUGGGGACAUUUCGAAGAUGC----------------CAAGAUGCGCCACCAGGAGCGAGUA .......(((((((.(((...((((.((((......))))(((((----((((...((((.....)))))----------------))))))))))))...)))))))))) ( -31.20) >DroEre_CAF1 15545 99 - 1 UUUGAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCGUU----UUGGGGACAUUUCGAAGAUGCUAAGUUGCGA--------AGAUGCGCCACCAGGAGAGAGUA .......(((((..((((...((((.((((......))))(((((----((.(.(((................))).).)--------))))))))))...)))).))))) ( -26.69) >DroYak_CAF1 14914 107 - 1 UUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCGUU----UUGUAGACAUUUCGAAGAUGCGAAGAUGUGAAGACGCCAAGAUGCGCCACCAGGAGCGAGUA .......(((((((.(((...((((......((....))((((((----((....((((((((......)).)))))).)))))))).......))))...)))))))))) ( -31.10) >DroAna_CAF1 18511 82 - 1 ----AGCUAUUUGC-UCUAUUUGCCACUUAUUCAAUUGAGCCGUUACAUUUGGGGACACUUUU-----GC-------------------GCUGCACCACCAGGAGCGAGUA ----...(((((((-(((...(((..((((......)))).(((......((....)).....-----))-------------------)..)))......)))))))))) ( -16.70) >consensus UUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUU____UUGGGGACAUUUCGAAGAUGC________________CAAGAUGCGCCACCAGGAGCGAGUA .......(((((((.(((...((((.((((......))))(((((.....((....)).......)))))........................))))...)))))))))) (-18.94 = -19.80 + 0.86)

| Location | 14,756,437 – 14,756,537 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 93.27 |
| Mean single sequence MFE | -28.78 |
| Consensus MFE | -25.56 |
| Energy contribution | -25.60 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.89 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.525529 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
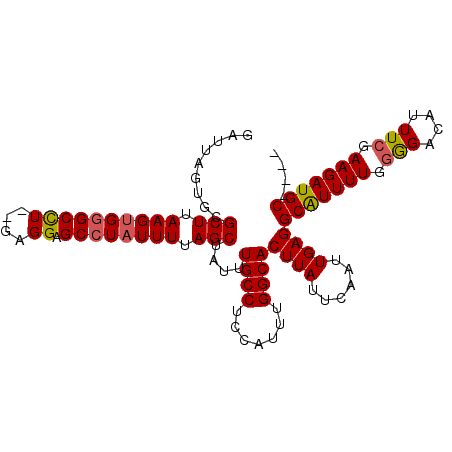
>3R_DroMel_CAF1 14756437 100 - 27905053 GAUUAGUGCGCUUAAGUGCGCUUAAGAGGAGCCU-UUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUUUUGGGGACAUUUCGAAGAUGC---- .......((((....))))(((.(((((....))-))).)))....((((.......))))((((......))))(((((((.(((....))).)))))))---- ( -26.60) >DroSec_CAF1 15396 99 - 1 GAUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUUUUGGGGACAUUUCGAAGAUGC---- .........(((.(((((((((.--...).)))))))).)))....((((.......))))((((......))))(((((((.(((....))).)))))))---- ( -27.70) >DroSim_CAF1 16557 99 - 1 GAUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUUUUGGGGACAUUUCGAAGAUGC---- .........(((.(((((((((.--...).)))))))).)))....((((.......))))((((......))))(((((((.(((....))).)))))))---- ( -27.70) >DroEre_CAF1 15574 103 - 1 GGUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUGAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCGUUUUGGGGACAUUUCGAAGAUGCUAAG (((.((((.(((((((((((((.--...).)))))))))))))))).))).....((((((((((......)))).((...((....))...))....)))))). ( -34.70) >DroYak_CAF1 14951 103 - 1 GGUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCGUUUUGUAGACAUUUCGAAGAUGCGAAG (((.((((.(((.(((((((((.--...).)))))))).))))))).)))(((((((.((.((((......))))))(((.....))).......))))).)).. ( -27.20) >consensus GAUUAGUGCGCUUAAGUGGGCCU__GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCAAUUGAGGCAUUUUGGGGACAUUUCGAAGAUGC____ .........(((.((((((((((....)).)))))))).)))....((((.......))))((((......))))(((((((.(((....))).))))))).... (-25.56 = -25.60 + 0.04)


| Location | 14,756,469 – 14,756,560 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 95.59 |
| Mean single sequence MFE | -25.28 |
| Consensus MFE | -22.26 |
| Energy contribution | -22.14 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.88 |
| SVM RNA-class probability | 0.981314 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14756469 91 + 27905053 UGAAUAAGUGCCAAAUGGAGGCAAAUAGCUAAAA-AGGCUCCUCUUAAGCGCACUUAAGCGCACUAAUCCAGUGCGACUGUGUACUCCGUAC ....(((((((.....(((((.....((((....-.))))))))).....)))))))((((((((.....)))))).))............. ( -27.10) >DroSec_CAF1 15428 90 + 1 UGAAUAAGUGCCAAAUGGAGGCAAAUAGCUAAAAUAGGCUCCUC--AGGCCCACUUAAGCGCACUAAUCCAGUGCGACUGUGUGCUCCGUAC .......((((......((((.....((((......))))))))--.((..(((...((((((((.....)))))).))..)))..)))))) ( -26.00) >DroSim_CAF1 16589 90 + 1 UGAAUAAGUGCCAAAUGGAGGCAAAUAGCUAAAAUAGGCUCCUC--AGGCCCACUUAAGCGCACUAAUCCAGUGCGACUGUGUGCUCCGUAC .......((((......((((.....((((......))))))))--.((..(((...((((((((.....)))))).))..)))..)))))) ( -26.00) >DroEre_CAF1 15610 90 + 1 UGAAUAAGUGCCAAAUGGAGGCAAAUAGCUCAAAUAGGCUCCUC--AGGCCCACUUAAGCGCACUAACCCAGUGCGACUGUGUACUCCAUAC ..............((((((....((((.((.....((((....--.)))).........(((((.....)))))))))))...)))))).. ( -23.90) >DroYak_CAF1 14987 90 + 1 UGAAUAAGUGCCAAAUGGAGGCAAAUAGCUAAAAUAGGCUCCUC--AGGCCCACUUAAGCGCACUAACCCAGUGCGACUGUGUACUCCGUAC ..............((((((((.....)).......((((....--.))))(((.....((((((.....))))))...)))..)))))).. ( -23.40) >consensus UGAAUAAGUGCCAAAUGGAGGCAAAUAGCUAAAAUAGGCUCCUC__AGGCCCACUUAAGCGCACUAAUCCAGUGCGACUGUGUACUCCGUAC ..............(((((((((....(((......((((.......))))......)))(((((.....))))).....))).)))))).. (-22.26 = -22.14 + -0.12)



| Location | 14,756,469 – 14,756,560 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 95.59 |
| Mean single sequence MFE | -28.14 |
| Consensus MFE | -24.48 |
| Energy contribution | -25.12 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.915249 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14756469 91 - 27905053 GUACGGAGUACACAGUCGCACUGGAUUAGUGCGCUUAAGUGCGCUUAAGAGGAGCCU-UUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCA ....(((((((..((.(((((((...)))))))))...))))(((.(((((....))-))).)))....((((.......))))....))). ( -26.20) >DroSec_CAF1 15428 90 - 1 GUACGGAGCACACAGUCGCACUGGAUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCA ....((((((.......((((((...))))))(((.(((((((((.--...).)))))))).)))....)).))))................ ( -27.30) >DroSim_CAF1 16589 90 - 1 GUACGGAGCACACAGUCGCACUGGAUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCA ....((((((.......((((((...))))))(((.(((((((((.--...).)))))))).)))....)).))))................ ( -27.30) >DroEre_CAF1 15610 90 - 1 GUAUGGAGUACACAGUCGCACUGGGUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUGAGCUAUUUGCCUCCAUUUGGCACUUAUUCA .....(((((.......((((((...))))))(((((((((((((.--...).))))))))))))....((((.......))))..))))). ( -32.00) >DroYak_CAF1 14987 90 - 1 GUACGGAGUACACAGUCGCACUGGGUUAGUGCGCUUAAGUGGGCCU--GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCA .....(((((.......((((((...))))))(((.(((((((((.--...).)))))))).)))....((((.......))))..))))). ( -27.90) >consensus GUACGGAGUACACAGUCGCACUGGAUUAGUGCGCUUAAGUGGGCCU__GAGGAGCCUAUUUUAGCUAUUUGCCUCCAUUUGGCACUUAUUCA .....(((((.......((((((...))))))(((.((((((((((....)).)))))))).)))....((((.......))))..))))). (-24.48 = -25.12 + 0.64)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:40 2006