
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,465,391 – 14,465,547 |
| Length | 156 |
| Max. P | 0.685262 |

| Location | 14,465,391 – 14,465,510 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.97 |
| Mean single sequence MFE | -45.83 |
| Consensus MFE | -38.44 |
| Energy contribution | -39.44 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685262 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14465391 119 - 27905053 AGGCGGAGGCGGGAUGCAGCCAGCUGCGUUGAGGCAACCCAACCGCCUGCCUUGAUGUGUCAAACCGCAAGCUGACAAAACCGCAAGCCAA-AACUGCGUUCUCAACGCAAUUGCAUUUU ..(((((((((((..((((....))))((((.(....).))))..))))))).....(((((...(....).)))))...))))..((...-((.(((((.....))))).))))..... ( -42.60) >DroSim_CAF1 23279 118 - 1 AGGCGGAGGCGGGAUGCAGCCAGCUGCGUUGAGGCAACCCAACCGCCUGCCUUGAUGUGUCAAG-CGCGGGCUGACAAAACCGCAAGCCAA-AACUGCGUUCUCAACGCAAUUGCAUUUU .(((....((((...((((....))))((((.(....).)))).(((((((((((....)))))-.))))))........))))..)))..-...(((((.....))))).......... ( -47.40) >DroYak_CAF1 25047 119 - 1 AGGCGGAGGCGGGAUGCAGCCAGCUGCGCUGAGGCAACCAAACCGCCUGCCCUGAUGUGUCAAG-CGCGGGCUGACAAAACCGCAAGCCAAAAACUGCGUUCUCAACGCAGUUGCAUUUU .(((....((((..(((((((....(((((.((((.........))))....(((....)))))-))).))))).))...))))..)))...((((((((.....))))))))....... ( -47.50) >consensus AGGCGGAGGCGGGAUGCAGCCAGCUGCGUUGAGGCAACCCAACCGCCUGCCUUGAUGUGUCAAG_CGCGGGCUGACAAAACCGCAAGCCAA_AACUGCGUUCUCAACGCAAUUGCAUUUU .(((....((((..(((((((.((((((((((((((...........)))))))))))).......)).))))).))...))))..)))......(((((.....))))).......... (-38.44 = -39.44 + 1.00)



| Location | 14,465,430 – 14,465,547 |
|---|---|
| Length | 117 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.11 |
| Mean single sequence MFE | -45.53 |
| Consensus MFE | -35.59 |
| Energy contribution | -35.60 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.603403 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14465430 117 - 27905053 ---GAUAGCGAGGACCCACUGUACUUUAGUGCCUCAUUUCAGGCGGAGGCGGGAUGCAGCCAGCUGCGUUGAGGCAACCCAACCGCCUGCCUUGAUGUGUCAAACCGCAAGCUGACAAAA ---..((((((((...(((((.....))))))))).......((((.((((((..((((....))))((((.(....).))))..))))))((((....)))).))))..))))...... ( -45.50) >DroSim_CAF1 23318 108 - 1 ---CAUAGCGAGCUUCCACUGUACUUCAG--------UUCAGGCGGAGGCGGGAUGCAGCCAGCUGCGUUGAGGCAACCCAACCGCCUGCCUUGAUGUGUCAAG-CGCGGGCUGACAAAA ---........((((((.(((.((....)--------).)))..))))))(((((((((....)))))))(.(....).)..))(((((((((((....)))))-.))))))........ ( -43.90) >DroYak_CAF1 25087 119 - 1 UGCCACACCGAGGCUCCACUGUACUUCAGCGCCUCAGUUCAGGCGGAGGCGGGAUGCAGCCAGCUGCGCUGAGGCAACCAAACCGCCUGCCCUGAUGUGUCAAG-CGCGGGCUGACAAAA .(((.....(((((....(((.....))).)))))...(((((.(.((((((...((((....))))((....)).......)))))).))))))((((.....-)))))))........ ( -47.20) >consensus ___CAUAGCGAGGAUCCACUGUACUUCAG_GCCUCA_UUCAGGCGGAGGCGGGAUGCAGCCAGCUGCGUUGAGGCAACCCAACCGCCUGCCUUGAUGUGUCAAG_CGCGGGCUGACAAAA ....((((.(......).))))...(((((........(((((.(.(((((((((((((....)))))))..(....)....)))))).))))))((((......)))).)))))..... (-35.59 = -35.60 + 0.01)

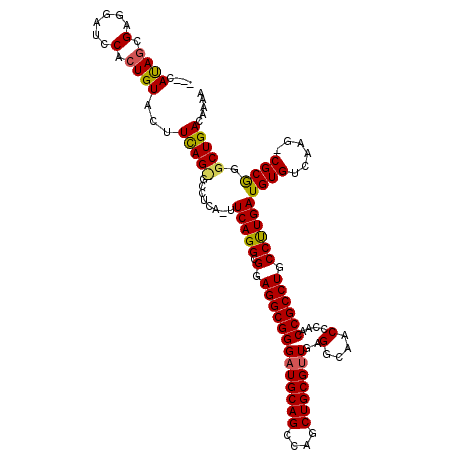

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:36 2006