
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,274,070 – 14,274,201 |
| Length | 131 |
| Max. P | 0.853991 |

| Location | 14,274,070 – 14,274,181 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 79.64 |
| Mean single sequence MFE | -19.14 |
| Consensus MFE | -13.92 |
| Energy contribution | -13.70 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853991 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14274070 111 - 27905053 GCUUAUGGGUCGUAAAGUUGACCUUCAAACAGCAACAUUUGCCGAAAAAAUACGAGCAACCAACACACACACACA----CGUGCGAACGAAUGUGCAAAAAAAAAACAAGACUAA (((...((((((......))))))......))).....((((((........)).))))..........((((..----(((....)))..)))).................... ( -19.50) >DroPse_CAF1 57854 87 - 1 GCUUAUGGGUCGUAAAGUUGACCUUCAAACAGCAACAUUUGCCGAAAAAAUACGAGCAGGCAGC---------CA----C---------------CAAAAAGAAAAAGAGACGAA (((...((((((......))))))......))).....(((((...............))))).---------..----.---------------.................... ( -14.56) >DroSim_CAF1 44753 114 - 1 GCUUAUGGGUCGUAAAGUUGACCUUCAAACAGCAACAUUUGCCGAAAAAAUACGAGCAACCAACACACACAGACACACACGUGCAAACGUAUGUGC-AAAAAAAAACACGACUAA (((...((((((......))))))......))).....((((((........)).))))............(.((((.((((....)))).)))))-.................. ( -22.50) >DroEre_CAF1 55422 108 - 1 GCUUAUGGGUCGUAAAGUUGACCUUCAAACAGCAACAUUUGCCGAAAAAAUACGAGCAACCGAAACACGCACACA----CGUGCACACGUAUGUGCAAAA---AAACACGACUAA (((...((((((......))))))......))).....((((((........)).)))).........(((((.(----(((....)))).)))))....---............ ( -26.20) >DroYak_CAF1 50173 106 - 1 GCUUAUGGGUCGUAAAGUUGACCUUCAAACAGCAACAUUUGCCGAAAAAAUACGAGCAACCAAAACACACACAC--------GCAUACG-AUGUGCAAAAAAAAAACACGACUAA (((...((((((......))))))......))).....((((((........)).))))...............--------((((...-..))))................... ( -17.50) >DroPer_CAF1 58184 87 - 1 GCUUAUGGGUCGUAAAGUUGACCUUCAAACAGCAACAUUUGCCGAAAAAAUACGAGCAGGCAGC---------CA----C---------------CAAAAAGAAAAAGAGACGAA (((...((((((......))))))......))).....(((((...............))))).---------..----.---------------.................... ( -14.56) >consensus GCUUAUGGGUCGUAAAGUUGACCUUCAAACAGCAACAUUUGCCGAAAAAAUACGAGCAACCAACACACACACACA____C__GCA_ACG_AUGUGCAAAAAAAAAACAAGACUAA (((...((((((......))))))......))).....((((((........)).))))........................................................ (-13.92 = -13.70 + -0.22)



| Location | 14,274,106 – 14,274,201 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 75.16 |
| Mean single sequence MFE | -17.49 |
| Consensus MFE | -11.02 |
| Energy contribution | -11.35 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.766506 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14274106 95 - 27905053 ----CA-AAAACACACA-------------UU-U-UUGGCGCUUAUGGGUCGUAAAGUUGACCUUCAAACA-GCAACAUUUGCCGAAA---AAAUACGAGCAACCAACACACACACACA ----..-..........-------------..-.-((((.(((...((((((......))))))......)-)).....((((((...---.....)).))))))))............ ( -16.80) >DroPse_CAF1 57875 103 - 1 ACAACA-CACACACACAUGAGCACAAUUUUUU-U-UUUGCGCUUAUGGGUCGUAAAGUUGACCUUCAAACA-GCAACAUUUGCCGAAA---AAAUACGAGCAGGCAGC---------CA ......-........(((((((.(((......-.-.))).)))))))(((((......)))))........-.......(((((....---...........))))).---------.. ( -22.76) >DroEre_CAF1 55455 95 - 1 ----CG-AAAACACACA-------------UU-U-UUGGCGCUUAUGGGUCGUAAAGUUGACCUUCAAACA-GCAACAUUUGCCGAAA---AAAUACGAGCAACCGAAACACGCACACA ----..-..........-------------.(-(-((((.(((...((((((......))))))......)-)).....((((((...---.....)).)))))))))).......... ( -18.30) >DroWil_CAF1 55556 87 - 1 ----CA-AAAACAUUUU-------------UUUU-UUUUUGCCUUUGGGUCGUA-AGUUGACCUUCAAACAAGCAACAUUUGCCGAAA---AAAUACGAACAUACACA---------CA ----..-.....(((((-------------(((.-...(((.....((((((..-...)))))).....)))(((.....))).))))---)))).............---------.. ( -11.80) >DroAna_CAF1 45742 100 - 1 ----CGCAAAACACACA-------------UU-UUUUUGCGCUUAUGGGUCGUAAAGUUGACCUUCAAACA-GCAACAUUUGCCGAAAAAAAAAUACGAGCAACCAAAAAACACACUCA ----.((((((......-------------..-.))))))((((..((((((......)))))).....(.-(((.....))).)............)))).................. ( -17.00) >DroPer_CAF1 58205 99 - 1 A------CACACACACAUGAGCACAAUUUUUUUU-UUUUCGCUUAUGGGUCGUAAAGUUGACCUUCAAACA-GCAACAUUUGCCGAAA---AAAUACGAGCAGGCAGC---------CA .------........(((((((............-.....)))))))(((((......)))))........-.......(((((....---...........))))).---------.. ( -18.29) >consensus ____CA_AAAACACACA_____________UU_U_UUUGCGCUUAUGGGUCGUAAAGUUGACCUUCAAACA_GCAACAUUUGCCGAAA___AAAUACGAGCAACCAAA_________CA ........................................((((..((((((......))))))........(((.....)))..............)))).................. (-11.02 = -11.35 + 0.33)

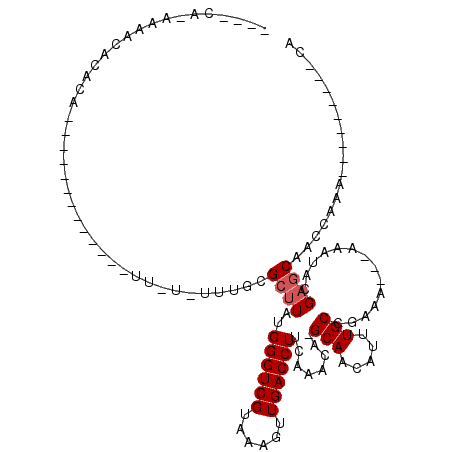

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:54:09 2006