
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,214,322 – 14,214,442 |
| Length | 120 |
| Max. P | 0.662313 |

| Location | 14,214,322 – 14,214,442 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.17 |
| Mean single sequence MFE | -39.32 |
| Consensus MFE | -22.11 |
| Energy contribution | -24.83 |
| Covariance contribution | 2.72 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.56 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.530500 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
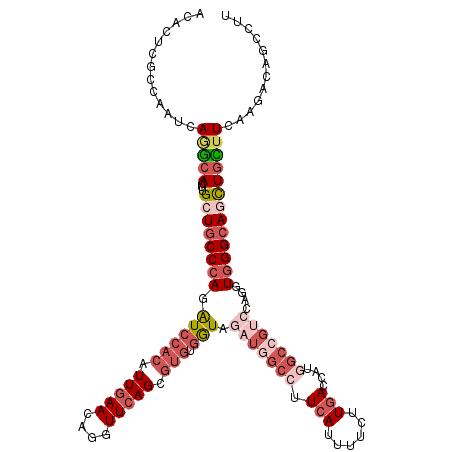
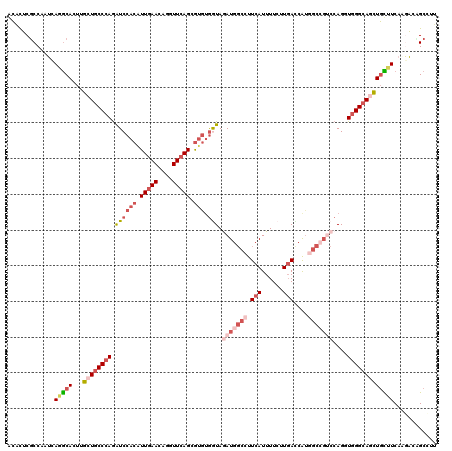
>3R_DroMel_CAF1 14214322 120 + 27905053 ACCUUCGCCAAUCAGGCACUUGCUGCCCAGAUCCACGUUGAAAAGGUUCAGCGUGUGGUAGAUGCCCUUCAUUUUCUUGACCAUGGCGCUCCAGGUGGGCAGCUGUUUCAAGGCAGCCUG ............(((((....((((((((....(((((((((....)))))))))(((.((.((((..(((......)))....)))))))))..))))))))((((....))))))))) ( -46.30) >DroVir_CAF1 93320 120 + 1 GCACUCGCCAAUCAAGCACUUGCUGCCCAGAUCCACAUUGAACAUGUUCAGCGUGUGAUAGAUGGCCUUCAUUUUCUUGACCAUGGCCGUCCAGUUGGGCAGCUGCUUCUUGAUUGCCUU ......(.((((((((.....(((((((((((((((.(((((....))))).))).))).(((((((.(((......)))....)))))))...))))))))).....)))))))).).. ( -48.90) >DroGri_CAF1 78793 120 + 1 GCACUCGCCAAUCAAGCACUUGCUGCCCAGGUCCACAUUGAACAUGUUCAGGGUGAAGUAAAUGGCCUUCAUUUUCUUGAUCAUGGCCGACCAGGUGGGCAGCUGCUUCAGAAUCGCCUU ......((...(((((((...(((((((((((((((.(((((....))))).)))........((((.(((......)))....))))))))...)))))))))))))..))...))... ( -41.90) >DroWil_CAF1 82022 120 + 1 AGCCUCGCCAAUUAGACAUUUGCUGCCCAGAUCCACAUUGAACAAGUUGAGGACAUGGUAAAUGGCCUUCAUUUUCUUCACCAUAGCCGACCAGGUGGGCAACUUUUUCAAAACCGCCUG .((...((.(((.....))).))(((((..........((((.(((.(((((.(((.....)))..))))).))).)))).....(((.....))))))))..............))... ( -23.00) >DroMoj_CAF1 98530 120 + 1 GCACUCGCCGAUCAUGCACUUGCUUCCCAGAUCCACAUUGAACAGGUUCAGCGUGUGGUAGAUGGCCUUCAUUUUCUUGACCAUGACCGUCCAGUUGGGCAGCUGCUUCUUGAUGGCCUU ......(((.((((.(((...(((.(((((..((((((((((....))))..))))))..(((((.(.(((......)))....).)))))...))))).))))))....)))))))... ( -35.80) >DroAna_CAF1 78481 120 + 1 ACCCUCUCCAAUGAGGCACUUGCUGCCCAAGUCCACAUUGAAAAGGUUCAGGGUGUGAUAGAUGGCCUUCAUUUUCUUGAUCGAAGCAGUCCAGUUCGGCAAUUGCUUCAGGGCAGCUUU ..((((......)))).....(((((((................(((.(((((.((((.((.....))))))..))))))))((((((((((.....))..)))))))).)))))))... ( -40.00) >consensus ACACUCGCCAAUCAGGCACUUGCUGCCCAGAUCCACAUUGAACAGGUUCAGCGUGUGGUAGAUGGCCUUCAUUUUCUUGACCAUGGCCGUCCAGGUGGGCAGCUGCUUCAAGACAGCCUU .............(((((...((((((((.((((((.(((((....))))).))).))).(((((((.(((......)))....)))))))....)))))))))))))............ (-22.11 = -24.83 + 2.72)

| Location | 14,214,322 – 14,214,442 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.17 |
| Mean single sequence MFE | -40.15 |
| Consensus MFE | -25.48 |
| Energy contribution | -26.10 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.662313 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
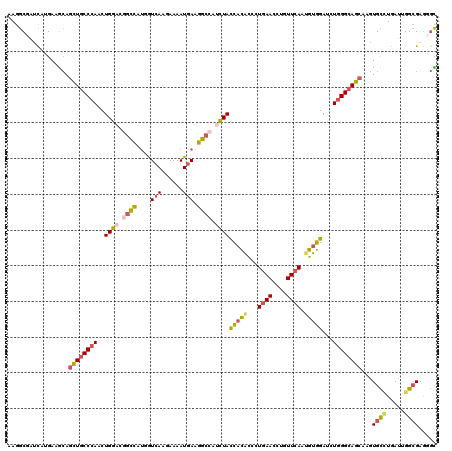
>3R_DroMel_CAF1 14214322 120 - 27905053 CAGGCUGCCUUGAAACAGCUGCCCACCUGGAGCGCCAUGGUCAAGAAAAUGAAGGGCAUCUACCACACGCUGAACCUUUUCAACGUGGAUCUGGGCAGCAAGUGCCUGAUUGGCGAAGGU ......(((((......((((((((..((((..(((....(((......)))..))).))))...((((.((((....)))).))))....))))))))...((((.....))))))))) ( -42.80) >DroVir_CAF1 93320 120 - 1 AAGGCAAUCAAGAAGCAGCUGCCCAACUGGACGGCCAUGGUCAAGAAAAUGAAGGCCAUCUAUCACACGCUGAACAUGUUCAAUGUGGAUCUGGGCAGCAAGUGCUUGAUUGGCGAGUGC ..(.((((((((.....((((((((..((((.((((....(((......))).)))).))))((.((((.((((....)))).))))))..)))))))).....)))))))).)...... ( -48.40) >DroGri_CAF1 78793 120 - 1 AAGGCGAUUCUGAAGCAGCUGCCCACCUGGUCGGCCAUGAUCAAGAAAAUGAAGGCCAUUUACUUCACCCUGAACAUGUUCAAUGUGGACCUGGGCAGCAAGUGCUUGAUUGGCGAGUGC ..(.(((((...(((((((((((((...((((((((....(((......))).))))........(((..((((....))))..)))))))))))))))...)))))))))).)...... ( -43.20) >DroWil_CAF1 82022 120 - 1 CAGGCGGUUUUGAAAAAGUUGCCCACCUGGUCGGCUAUGGUGAAGAAAAUGAAGGCCAUUUACCAUGUCCUCAACUUGUUCAAUGUGGAUCUGGGCAGCAAAUGUCUAAUUGGCGAGGCU .....((((((......((((((((...((((.((.....((((.((..(((.((.(((.....))).)))))..)).))))..)).))))))))))))....(((.....))))))))) ( -34.00) >DroMoj_CAF1 98530 120 - 1 AAGGCCAUCAAGAAGCAGCUGCCCAACUGGACGGUCAUGGUCAAGAAAAUGAAGGCCAUCUACCACACGCUGAACCUGUUCAAUGUGGAUCUGGGAAGCAAGUGCAUGAUCGGCGAGUGC ...(((((((....((((((.((((..((((.((((....(((......))).)))).))))(((((...((((....)))).)))))...)))).)))...))).)))).)))...... ( -35.80) >DroAna_CAF1 78481 120 - 1 AAAGCUGCCCUGAAGCAAUUGCCGAACUGGACUGCUUCGAUCAAGAAAAUGAAGGCCAUCUAUCACACCCUGAACCUUUUCAAUGUGGACUUGGGCAGCAAGUGCCUCAUUGGAGAGGGU ..((.((.(((((((((....((.....))..))))))..(((......)))))).)).)).............(((((((((((.(((((((.....))))).)).))))))))))).. ( -36.70) >consensus AAGGCGAUCAUGAAGCAGCUGCCCAACUGGACGGCCAUGGUCAAGAAAAUGAAGGCCAUCUACCACACCCUGAACCUGUUCAAUGUGGAUCUGGGCAGCAAGUGCCUGAUUGGCGAGGGC .................((((((((..((((.((((....(((......))).)))).))))(((((...((((....)))).)))))...))))))))...((((.....))))..... (-25.48 = -26.10 + 0.62)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:53 2006