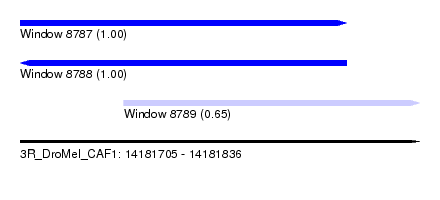

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,181,705 – 14,181,836 |
| Length | 131 |
| Max. P | 0.999612 |

| Location | 14,181,705 – 14,181,812 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 84.74 |
| Mean single sequence MFE | -42.40 |
| Consensus MFE | -35.72 |
| Energy contribution | -35.60 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.95 |
| Structure conservation index | 0.84 |
| SVM decision value | 3.78 |
| SVM RNA-class probability | 0.999612 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
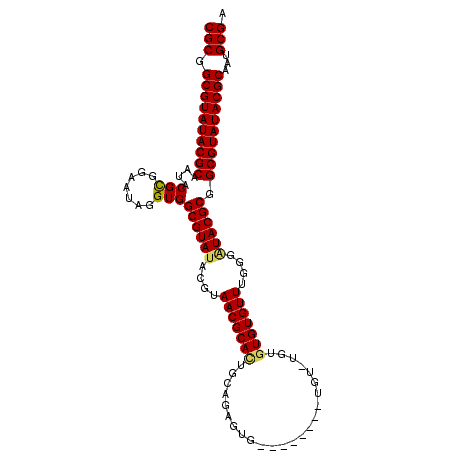
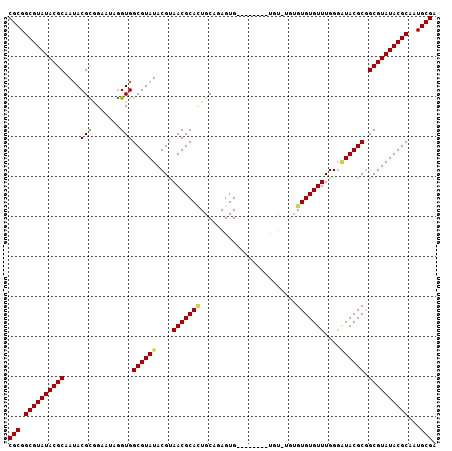
>3R_DroMel_CAF1 14181705 107 + 27905053 CGCGGCGUAUACGCAAUACGCGGAAUAGGUGGCGUAUACGUAACGCAAUGCAGAGUGAAAAUGUGUGUAUGUGUGUGUUUGGAAUACGCGGCGUAUACGCAAUGCGA (((.((((((((((....(((.......)))((((((.((..((((((((((..((....))...))))).)))))...))..)))))).))))))))))...))). ( -41.40) >DroSec_CAF1 173916 103 + 1 CGCGGCGUAUACGCAAAACGCGGAAUAGGUGGCGUAUACGUAACGCACUGCAGAGUGUAU----AUGUGUGUGUGUGUUUGGGUUACGCGGCGUAUACGCAAUGCGA .(((((((((((((...((.(......))).))))))))))..)))..((((..((((((----((((.((((((.........))))))))))))))))..)))). ( -42.10) >DroSim_CAF1 59213 105 + 1 CGCGGCGUAUACGCAAUACGCGGAAUAGGUGGCGUAUACGCAACGCACUGCAGAGUGUGU--GUCUGUGUGUGUGUGUUUGGGAUACGCGGCGUAUACGCAAUGCGA (((.((((((((((.(((((((((.......(((....))).(((((((....)))))))--.)))))))))(((((((...))))))).))))))))))...))). ( -50.10) >DroEre_CAF1 51759 93 + 1 CGCGGCGUAUACGCAAUACGUGGCAUAGGUGGCGUAUACGUAACGCACUGCACAGUG--------------GGUGUGUUUGGGGUACGCGGCGUAUACGCAAUGCGA (((.((((((((((.((((((.((....)).)))))).((((.(.((..(((((...--------------..))))).)).).))))..))))))))))...))). ( -41.00) >DroYak_CAF1 37045 93 + 1 CGCGGCGUAUACGCAAUACGUAGGAUGAGUGGCGUAUACGUAACGCAUUGCGGAGUG--------------UGUGUGUUUGGGGUACGCAGCGUAUACGCAAUGCGA (((.((((((((((..(((.........)))((((((.((..((((((........)--------------)))))...))..)))))).))))))))))...))). ( -37.40) >consensus CGCGGCGUAUACGCAAUACGCGGAAUAGGUGGCGUAUACGUAACGCACUGCAGAGUG________UGU_UGUGUGUGUUUGGGAUACGCGGCGUAUACGCAAUGCGA (((.((((((((((....(((.......)))((((((....(((((((........................)))))))....)))))).))))))))))...))). (-35.72 = -35.60 + -0.12)

| Location | 14,181,705 – 14,181,812 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 84.74 |
| Mean single sequence MFE | -28.74 |
| Consensus MFE | -24.14 |
| Energy contribution | -24.30 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.80 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.69 |
| SVM RNA-class probability | 0.996406 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
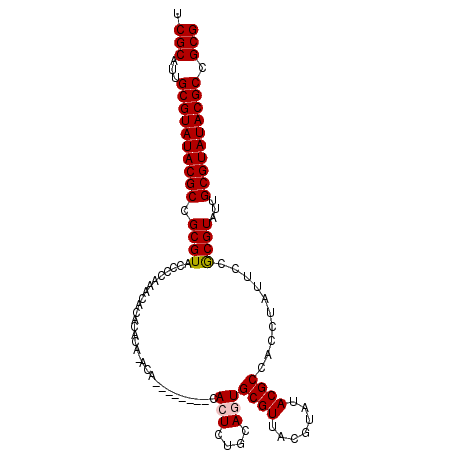
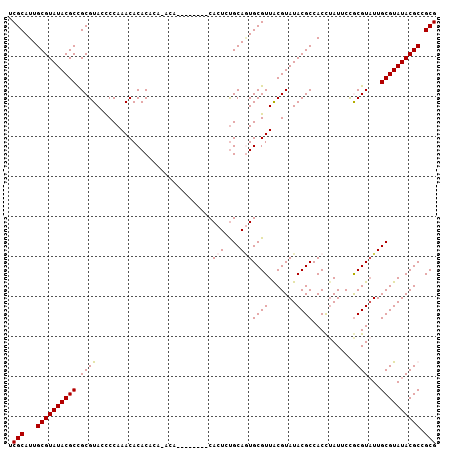
>3R_DroMel_CAF1 14181705 107 - 27905053 UCGCAUUGCGUAUACGCCGCGUAUUCCAAACACACACAUACACACAUUUUCACUCUGCAUUGCGUUACGUAUACGCCACCUAUUCCGCGUAUUGCGUAUACGCCGCG .(((...(((((((((..(((((.....................................)))))..)))))))))..........((((((.....)))))).))) ( -26.79) >DroSec_CAF1 173916 103 - 1 UCGCAUUGCGUAUACGCCGCGUAACCCAAACACACACACACAU----AUACACUCUGCAGUGCGUUACGUAUACGCCACCUAUUCCGCGUUUUGCGUAUACGCCGCG .(((...((((((((((.(((((((.((..((...........----........))...)).)))))))..((((..........))))...)))))))))).))) ( -28.11) >DroSim_CAF1 59213 105 - 1 UCGCAUUGCGUAUACGCCGCGUAUCCCAAACACACACACACAGAC--ACACACUCUGCAGUGCGUUGCGUAUACGCCACCUAUUCCGCGUAUUGCGUAUACGCCGCG .(((...((((((((((.((((((................((((.--......))))..)))))).))))))))))..........((((((.....)))))).))) ( -33.27) >DroEre_CAF1 51759 93 - 1 UCGCAUUGCGUAUACGCCGCGUACCCCAAACACACC--------------CACUGUGCAGUGCGUUACGUAUACGCCACCUAUGCCACGUAUUGCGUAUACGCCGCG ..((((.(((((((((..((((((......((((..--------------...))))..))))))..))))))))).....))))..(((...(((....))).))) ( -29.50) >DroYak_CAF1 37045 93 - 1 UCGCAUUGCGUAUACGCUGCGUACCCCAAACACACA--------------CACUCCGCAAUGCGUUACGUAUACGCCACUCAUCCUACGUAUUGCGUAUACGCCGCG ..((((((((.......((.((.......)).))..--------------.....))))))))((..(((((((((....(.......)....)))))))))..)). ( -26.04) >consensus UCGCAUUGCGUAUACGCCGCGUACCCCAAACACACACA_ACA________CACUCUGCAGUGCGUUACGUAUACGCCACCUAUUCCGCGUAUUGCGUAUACGCCGCG .(((...((((((((((.((((.............................(((....)))((((.......))))..........))))...)))))))))).))) (-24.14 = -24.30 + 0.16)

| Location | 14,181,739 – 14,181,836 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 80.36 |
| Mean single sequence MFE | -25.32 |
| Consensus MFE | -15.54 |
| Energy contribution | -15.14 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.651818 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14181739 97 + 27905053 UAUACGUAACGCAAUGCAGAGUGAAAAUGUGUGUAUGUGUGUGUUUGGAAUACGCGGCGUAUACGCAAUGCGAACGGGGACAAAAAGAAGAGACAGA .....((..((((.(((...((....))..((((((((((((((.....)))))).))))))))))).))))...(....)...........))... ( -25.60) >DroSec_CAF1 173950 93 + 1 UAUACGUAACGCACUGCAGAGUGUAU----AUGUGUGUGUGUGUUUGGGUUACGCGGCGUAUACGCAAUGCGAGCGGGUACAAAAAGAAGAGACAGA .....(((.(((..((((..((((((----((((.((((((.........))))))))))))))))..)))).)))..)))................ ( -28.20) >DroSim_CAF1 59247 95 + 1 UAUACGCAACGCACUGCAGAGUGUGU--GUCUGUGUGUGUGUGUUUGGGAUACGCGGCGUAUACGCAAUGCGAGCGGGGACAAAAAGAAGAGACAGA ((((((((((((((........))))--)).))))))))(((.(((......(((.(((....)))...)))...(....).......))).))).. ( -28.30) >DroEre_CAF1 51793 73 + 1 UAUACGUAACGCACUGCACAGUG--------------GGUGUGUUUGGGGUACGCGGCGUAUACGCAAUGCGAGCGGGGACAAAAAG---------- ....((.((((((((........--------------))))))))))..((.(((.(((....)))...))).))(....)......---------- ( -20.00) >DroYak_CAF1 37079 83 + 1 UAUACGUAACGCAUUGCGGAGUG--------------UGUGUGUUUGGGGUACGCAGCGUAUACGCAAUGCGAGCGGGGACAAAAGGAAGUGACAGA ..(((....(((((((((..(((--------------(((((((.......))))).))))).)))))))))...(....)........)))..... ( -24.50) >consensus UAUACGUAACGCACUGCAGAGUG________UGU_UGUGUGUGUUUGGGAUACGCGGCGUAUACGCAAUGCGAGCGGGGACAAAAAGAAGAGACAGA ....((((((((((........................))))))).......(((((((....)))..)))).)))..................... (-15.54 = -15.14 + -0.40)

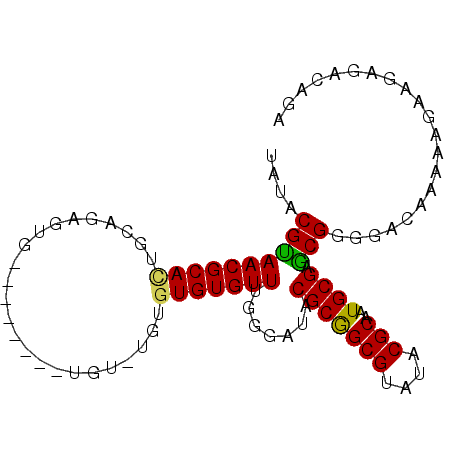
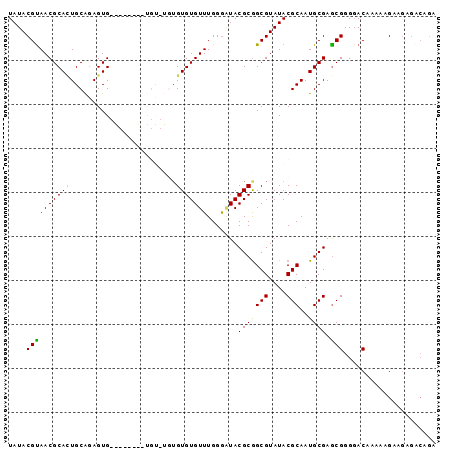
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:53:43 2006