
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 13,893,428 – 13,893,546 |
| Length | 118 |
| Max. P | 0.912187 |

| Location | 13,893,428 – 13,893,546 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.06 |
| Mean single sequence MFE | -24.13 |
| Consensus MFE | -19.38 |
| Energy contribution | -19.72 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.912187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13893428 118 + 27905053 -CAUU-CCAUUGCCCCCUUUUUUUGCUGCCCCGUUAAUAAGCGUAGAUUAUUUAUGUUUAUUAACCCUUAACAUGUUUACAUUUUGUAAAACGAGACUCUUAAUGUGAACUUUAUCAGGG -....-........((((..............(((((((((((((((...))))))))))))))).........(((((((((..((........))....)))))))))......)))) ( -24.70) >DroPse_CAF1 227162 108 + 1 -AGAGAAGAGU-UCC------U---CUUGCCCGUUAAUAAGCGUAGAUUAUUUAUGUUUAUUAACCCUUAACAUGUUUACAUUUUGUAAAACGAGACUCUUAAUGUGAACUUUAUCGGC- -....(((((.-..)------)---)))(((.(((((((((((((((...))))))))))))))).........(((((((((..((........))....)))))))))......)))- ( -26.00) >DroGri_CAF1 193540 112 + 1 CUCUC-AAAGUGCCGU---CCUUU----GGCCGUUAAUAAGCGUAGAUUAUUUAUGUUUAUUAACCCUUAACAUGUUUACAUUUUGUAAAACGAGACUCUUAAUGUGAACUUUAUCAUGA .....-...(.((((.---....)----))))(((((((((((((((...))))))))))))))).........(((((((((..((........))....))))))))).......... ( -25.40) >DroWil_CAF1 147771 98 + 1 ---------------------UUGGAGUCGCCGUUAAUAAGCUGAGAUUAUUUAUGUUUAUUAACCCUUAACAUGUUUACAUUUUGUAAAACGAGACUCUUAAUGUGAACUUUAUCAUU- ---------------------..((((((...((((((((((((((....)))).))))))))))..........(((((.....)))))....))))))....((((......)))).- ( -19.10) >DroMoj_CAF1 225328 111 + 1 CUCUC-AAAACGCCGU---CCUUU----GGCCGUUAAUAAGCGUAGAUUAUUUAUGUUUAUUAACCCUUAACAUGUUUACAUU-UGUAAAACGAGACUCUUAAUGUGAACUUUAUCGUGA .....-.....((((.---....)----))).(((((((((((((((...))))))))))))))).........(((((((((-.((........))....))))))))).......... ( -24.80) >DroPer_CAF1 219767 108 + 1 -AGAGAAGAGU-CCC------U---CUUGCCCGUUAAUAAGCGUAGAUUAUUUAUGUUUAUUAACCCUUAACAUGUUUACAUUUUGUAAAACGAGACUCUUAAUGUGAACUUUAUCGGU- -....(((((.-..)------)---)))(((.(((((((((((((((...))))))))))))))).........(((((((((..((........))....)))))))))......)))- ( -24.80) >consensus _ACUC_AAAGU_CCC______UUU_CUUGCCCGUUAAUAAGCGUAGAUUAUUUAUGUUUAUUAACCCUUAACAUGUUUACAUUUUGUAAAACGAGACUCUUAAUGUGAACUUUAUCAGG_ ................................(((((((((((((((...))))))))))))))).........(((((((((..((........))....))))))))).......... (-19.38 = -19.72 + 0.33)

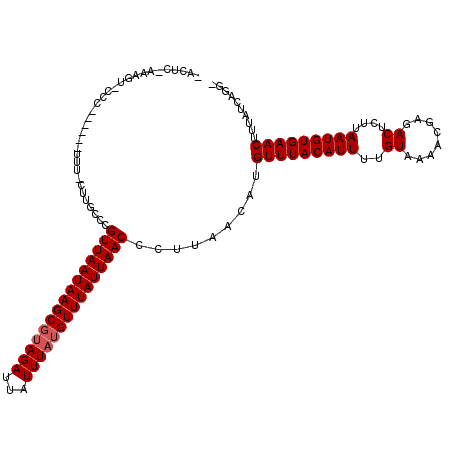

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:51:44 2006