
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 13,771,200 – 13,771,322 |
| Length | 122 |
| Max. P | 0.930583 |

| Location | 13,771,200 – 13,771,292 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 78.40 |
| Mean single sequence MFE | -21.36 |
| Consensus MFE | -15.11 |
| Energy contribution | -14.62 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.930583 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13771200 92 + 27905053 UCCCCGCUCUCCAUUUCCCCCCCGUU-UUAGUCGCAA---ACAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACG .........(((((((......((..-.....))...---..))))-----)))....(((((((((((((.....))))))))..))))).......... ( -22.20) >DroSec_CAF1 43751 93 + 1 UCCCGGCUCUCCAUUUCCCCCCGUUUUUCAGACCCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACG ....(((..(((((((......(((.....)))....---..))))-----)))....((..(((((((((.....)))))))))..)))))......... ( -24.00) >DroSim_CAF1 42641 93 + 1 UCCCGGCUCUCCAUUUCCCCCCGUUUUUCAGACCCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACG ....(((..(((((((......(((.....)))....---..))))-----)))....((..(((((((((.....)))))))))..)))))......... ( -24.00) >DroEre_CAF1 42458 85 + 1 -----GCUCUCCAUUUGCCCCC---UUGCAGCCCCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACG -----....((((((((.....---(((......)))---.)))))-----)))....(((((((((((((.....))))))))..))))).......... ( -23.70) >DroYak_CAF1 50134 90 + 1 UCCCGGCUCUCCAUUUCCCCCC---UUUCAGCAUCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACG ....(((..(((((((......---............---..))))-----)))....((..(((((((((.....)))))))))..)))))......... ( -21.75) >DroAna_CAF1 45198 87 + 1 UCUCUACUCUAC-------UCC---UUUCUCGCCCAAUGUCUAGCUUCUUUUUUUUUUUCAAUGAAAUUGCGUGAUGCAAUUGCAAUGUGGCAAGAU---- ............-------...---..(((.(((((.(((...((.((......((((.....))))......)).))....))).)).))).))).---- ( -12.50) >consensus UCCCGGCUCUCCAUUUCCCCCC___UUUCAGACCCAA___CCAGAU_____GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACG ....................................................((((..(((((((((((((.....)))))))))..)))).))))..... (-15.11 = -14.62 + -0.50)



| Location | 13,771,200 – 13,771,292 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 78.40 |
| Mean single sequence MFE | -25.37 |
| Consensus MFE | -15.88 |
| Energy contribution | -16.38 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.638296 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13771200 92 - 27905053 CGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGU---UUGCGACUAA-AACGGGGGGGAAAUGGAGAGCGGGGA (((....((.(((((..((((((((.....))))))))))))).)).(((-----.(((((---((.......)-))))))..)))........))).... ( -30.50) >DroSec_CAF1 43751 93 - 1 CGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGGGUCUGAAAAACGGGGGGAAAUGGAGAGCCGGGA .......(((((.((((((((((((.....)))))))).........(((-----.(((.(---((..........))).))))))...)))).))))).. ( -27.20) >DroSim_CAF1 42641 93 - 1 CGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGGGUCUGAAAAACGGGGGGAAAUGGAGAGCCGGGA .......(((((.((((((((((((.....)))))))).........(((-----.(((.(---((..........))).))))))...)))).))))).. ( -27.20) >DroEre_CAF1 42458 85 - 1 CGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGGGGCUGCAA---GGGGGCAAAUGGAGAGC----- ...........((.(((((..(((((((.((((.......))))...(((-----(.....---.)))))))))))---....)))))))......----- ( -24.90) >DroYak_CAF1 50134 90 - 1 CGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGAUGCUGAAA---GGGGGGAAAUGGAGAGCCGGGA .......(((((.((((((((((((.....)))))))).........(((-----..((..---((......))..---))..)))...)))).))))).. ( -25.10) >DroAna_CAF1 45198 87 - 1 ----AUCUUGCCACAUUGCAAUUGCAUCACGCAAUUUCAUUGAAAAAAAAAAAGAAGCUAGACAUUGGGCGAGAAA---GGA-------GUAGAGUAGAGA ----.(((((((.((.((.((((((.....)))))).)).))...............(........))))))))..---...-------............ ( -17.30) >consensus CGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC_____AUCUGG___UUGGGGCUGAAA___GGGGGGAAAUGGAGAGCCGGGA .......((.((((..(((((((((.....))))))))))))).))....................................................... (-15.88 = -16.38 + 0.50)



| Location | 13,771,226 – 13,771,322 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 80.61 |
| Mean single sequence MFE | -25.95 |
| Consensus MFE | -17.40 |
| Energy contribution | -18.35 |
| Covariance contribution | 0.95 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.809442 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13771226 96 + 27905053 UUAGUCGCAA---ACAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACGAGAAGAGGAAUUUCUUUCCAUUUUCGAGUU .........(---((...(-----(((.((.(((((((((((((.....))))))))..))))).)).))))((((((..((((.....)))).)))))).))) ( -27.30) >DroSec_CAF1 43778 96 + 1 UCAGACCCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACGAGCAGAGGAAUUUCUUUCCAUUUCCGAGUU ..........---.....(-----(((....(((((((((((((.....))))))))..)))))................((((.....))))...)))).... ( -24.40) >DroSim_CAF1 42668 96 + 1 UCAGACCCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACGAGUAGAGGAAUUUCUUUCCAUUUCCGAGUU ..........---.....(-----(((....(((((((((((((.....))))))))..)))))................((((.....))))...)))).... ( -24.40) >DroEre_CAF1 42477 96 + 1 GCAGCCCCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACGGGUAGAGGAAUUCCUUUCCAUUUCCCAGAU ...((((...---((....-----)).....(((((((((((((.....))))))))..))))).........))))...((((.....))))........... ( -28.20) >DroYak_CAF1 50158 96 + 1 UCAGCAUCAA---CCAGAU-----GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACGGGCAGAGGAAUUUCUUUCCAUUUCCCAGAU ....((((..---...)))-----)......(((((((((((((.....))))))))..))))).........(((.(((((((.....)))).)))))).... ( -29.50) >DroAna_CAF1 45215 94 + 1 UCUCGCCCAAUGUCUAGCUUCUUUUUUUUUUUCAAUGAAAUUGCGUGAUGCAAUUGCAAUGUGGCAAGAU----------GGAAUUUCUGCCAUCUUCCGGCUU ....(((.........................((.((.((((((.....)))))).)).)).((.(((((----------((........)))))))))))).. ( -21.90) >consensus UCAGACCCAA___CCAGAU_____GGAUUUCGCACUGCAAUUGCGUGCUGCAAUUGCAAAGUGCCAAUUUUACGAGCAGAGGAAUUUCUUUCCAUUUCCGAGUU ........................(((....(((((((((((((.....)))))))))..))))................((((.....))))...)))..... (-17.40 = -18.35 + 0.95)



| Location | 13,771,226 – 13,771,322 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 80.61 |
| Mean single sequence MFE | -28.20 |
| Consensus MFE | -17.00 |
| Energy contribution | -17.70 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.884288 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13771226 96 - 27905053 AACUCGAAAAUGGAAAGAAAUUCCUCUUCUCGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGU---UUGCGACUAA .....(((...((((.....))))..)))(((((((....(((((..((((((((.....)))))))))))))((.....-----.))..)---)))))).... ( -26.70) >DroSec_CAF1 43778 96 - 1 AACUCGGAAAUGGAAAGAAAUUCCUCUGCUCGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGGGUCUGA .((((((..(((((.(((......)))..........((.(((((..((((((((.....))))))))))))).)).)))-----))....---)))))).... ( -28.10) >DroSim_CAF1 42668 96 - 1 AACUCGGAAAUGGAAAGAAAUUCCUCUACUCGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGGGUCUGA .((((((..(((((.(((......)))..........((.(((((..((((((((.....))))))))))))).)).)))-----))....---)))))).... ( -28.10) >DroEre_CAF1 42477 96 - 1 AUCUGGGAAAUGGAAAGGAAUUCCUCUACCCGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGGGGCUGC ....(((....((((.....))))....)))......((.(((((..((((((((.....))))))))))))).)).(((-----(.....---.))))..... ( -30.00) >DroYak_CAF1 50158 96 - 1 AUCUGGGAAAUGGAAAGAAAUUCCUCUGCCCGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC-----AUCUGG---UUGAUGCUGA ....(((....((((.....))))....)))......((.(((((..((((((((.....))))))))))))).))...(-----(((...---..)))).... ( -30.60) >DroAna_CAF1 45215 94 - 1 AAGCCGGAAGAUGGCAGAAAUUCC----------AUCUUGCCACAUUGCAAUUGCAUCACGCAAUUUCAUUGAAAAAAAAAAAGAAGCUAGACAUUGGGCGAGA ..((((((((((((.(....).))----------))))).)).((.((.((((((.....)))))).)).)).........................))).... ( -25.70) >consensus AACUCGGAAAUGGAAAGAAAUUCCUCUACUCGUAAAAUUGGCACUUUGCAAUUGCAGCACGCAAUUGCAGUGCGAAAUCC_____AUCUGG___UUGGGGCUGA ...........((((.....)))).............((.((((..(((((((((.....))))))))))))).))............................ (-17.00 = -17.70 + 0.70)

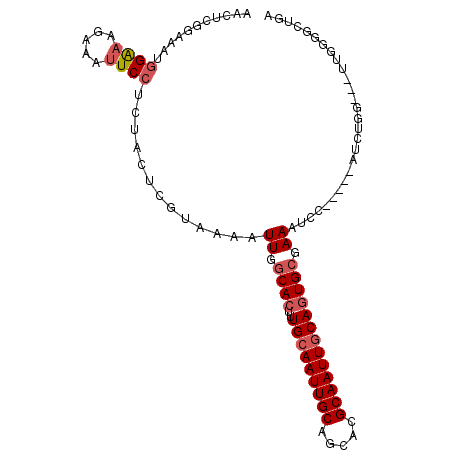

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:58 2006