
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 13,533,035 – 13,533,173 |
| Length | 138 |
| Max. P | 0.961804 |

| Location | 13,533,035 – 13,533,133 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 95.50 |
| Mean single sequence MFE | -24.07 |
| Consensus MFE | -22.15 |
| Energy contribution | -22.14 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.608463 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13533035 98 - 27905053 UAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACAUUCAUUGGAUGUUUGGCCGUGGGACCACCCAG----UAGUCCCCCAU ..((((((......)))))).......((((.((((((....(((................))))))))).))))(((((.((...)----).))))).... ( -24.49) >DroSec_CAF1 7769 102 - 1 UAUGUUAUUAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACAUUCAUUGGAUGUUUGGCCGUGGGACCACCCAGUAAGUAGUCCCCUAU ..(((((((....))))))).......((((.((((((....(((................))))))))).))))(((((.((.......)).))))).... ( -24.89) >DroEre_CAF1 7664 98 - 1 UAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCCAUUUGCACAUUCAUUGGAUGUUUGACCGUGGGACCACCCAG----UAGUCCCCCAU ..((((((......))))))((.....(((((.((.....)).)))))..))(((((.....)))))......(.(((((.((...)----).))))).).. ( -22.40) >DroYak_CAF1 8458 98 - 1 UAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACGUUCAUUGGAUGUUUGGCCGUGGGACCACCCAG----UAGUCCCCCAU ..((((((......)))))).......((((.((((((....(((................))))))))).))))(((((.((...)----).))))).... ( -24.49) >consensus UAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACAUUCAUUGGAUGUUUGGCCGUGGGACCACCCAG____UAGUCCCCCAU ..((((((......)))))).......((((.((((((....(((....((......))..))))))))).))))(((((.............))))).... (-22.15 = -22.14 + -0.00)


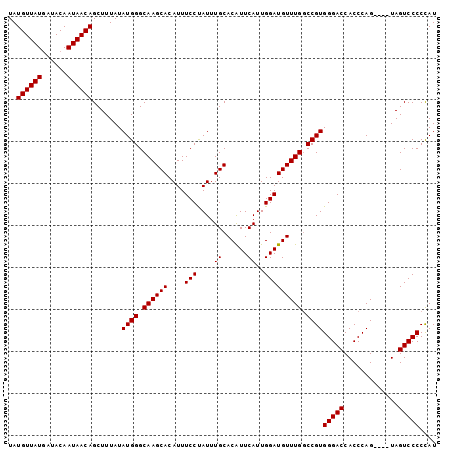
| Location | 13,533,056 – 13,533,173 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 94.00 |
| Mean single sequence MFE | -31.00 |
| Consensus MFE | -26.98 |
| Energy contribution | -29.23 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13533056 117 + 27905053 CCACGGCCAAACAUCCAAUGAAUGUGCAAAUAGGAAAUGUGCUUGCCCAUAUAAAGCUGUUAUUGUAUCAUAACAUACUAUGGGUGAAUGCAUUACCGCACAUUCGAUUGGAACGUA .............((((((((((((((.....((.(((((..(..((((((......((((((......))))))...))))))..)..))))).)))))))))).))))))..... ( -38.60) >DroSec_CAF1 7794 117 + 1 CCACGGCCAAACAUCCAAUGAAUGUGCAAAUAGGAAAUGUGCUUGCCCAUAUAAAGCUGUUAUUGUAUAAUAACAUAAUAUGUGUGAAUGCAUUACCGCACAUUCGAUUGGAGCUUA .............((((((((((((((.....((.(((((..(..(.(((((.....(((((((....)))))))..))))).)..)..))))).)))))))))).))))))..... ( -33.60) >DroEre_CAF1 7685 115 + 1 CCACGGUCAAACAUCCAAUGAAUGUGCAAAUGGGAAAUGUGCUUGCCCAUAUAAAGCUGUUAUUGUAUCAUAACAUACUAUGGCUGAUUGCAUUACCGC--AUUCGAUUGGAGCUUA .............(((((((((((((..((((.((.(((((......)))))..((((((...(((......)))....))))))..)).))))..)))--)))).))))))..... ( -26.30) >DroYak_CAF1 8479 115 + 1 CCACGGCCAAACAUCCAAUGAACGUGCAAAUAGGAAAUGUGCUUGCCCAUAUAAAGCUGUUAUUGUAUCAUAACAUACUACGGCUGAAUGCAUUACCGC--AUUCGAUUGGAGCUUA .((((..((.........))..))))..................(((((.....((((((...(((......)))....))))))((((((......))--))))...))).))... ( -25.50) >consensus CCACGGCCAAACAUCCAAUGAAUGUGCAAAUAGGAAAUGUGCUUGCCCAUAUAAAGCUGUUAUUGUAUCAUAACAUACUAUGGCUGAAUGCAUUACCGC__AUUCGAUUGGAGCUUA .............((((((((((((((.....((.(((((..(((((((((......((((((......))))))...)))))))))..))))).)))))))))).))))))..... (-26.98 = -29.23 + 2.25)

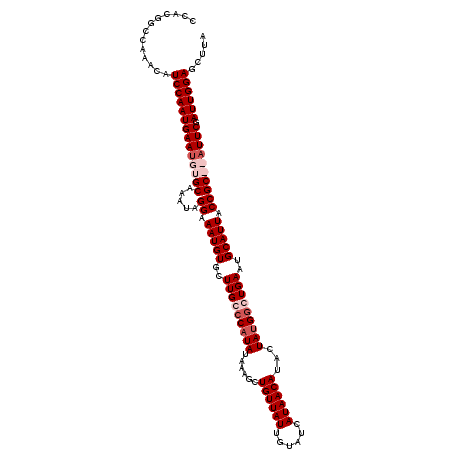
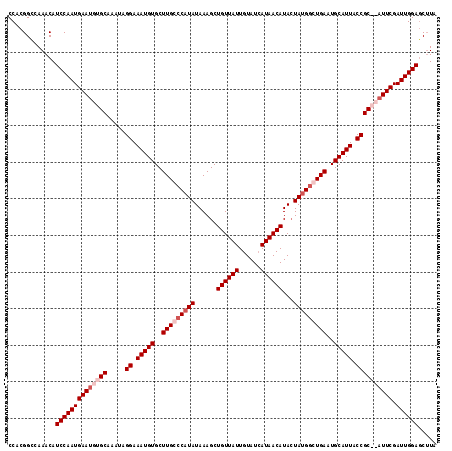
| Location | 13,533,056 – 13,533,173 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 94.00 |
| Mean single sequence MFE | -31.02 |
| Consensus MFE | -26.44 |
| Energy contribution | -27.88 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.832626 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13533056 117 - 27905053 UACGUUCCAAUCGAAUGUGCGGUAAUGCAUUCACCCAUAGUAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACAUUCAUUGGAUGUUUGGCCGUGG .((((.(((((.((((((((((.((((..((..((((((...((((((......))))))......))))))..))..)))).)).....))))))))))))))))).......... ( -34.80) >DroSec_CAF1 7794 117 - 1 UAAGCUCCAAUCGAAUGUGCGGUAAUGCAUUCACACAUAUUAUGUUAUUAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACAUUCAUUGGAUGUUUGGCCGUGG (((((((((((.((((((((((.((((........(((((..(((((((....))))))).....)))))........)))).)).....)))))))))))))).)))))....... ( -31.79) >DroEre_CAF1 7685 115 - 1 UAAGCUCCAAUCGAAU--GCGGUAAUGCAAUCAGCCAUAGUAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCCAUUUGCACAUUCAUUGGAUGUUUGACCGUGG (((((((((((.((((--(......(((((......((((..((((((......))))))...))))(((((.((.....)).))))))))))))))))))))).)))))....... ( -27.60) >DroYak_CAF1 8479 115 - 1 UAAGCUCCAAUCGAAU--GCGGUAAUGCAUUCAGCCGUAGUAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACGUUCAUUGGAUGUUUGGCCGUGG .((((.......((((--(((....)))))))..........((((((......))))))))))..(((((.((((((....(((................))))))))).))))). ( -29.89) >consensus UAAGCUCCAAUCGAAU__GCGGUAAUGCAUUCACCCAUAGUAUGUUAUGAUACAAUAACAGCUUUAUAUGGGCAAGCACAUUUCCUAUUUGCACAUUCAUUGGAUGUUUGGCCGUGG (((((((((((.((((((((((.((((......((((((...((((((......))))))......))))))......)))).)).....)))))))))))))).)))))....... (-26.44 = -27.88 + 1.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:47:29 2006