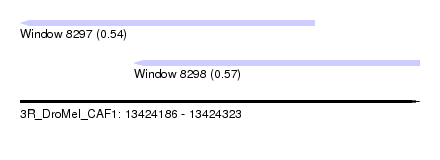
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 13,424,186 – 13,424,323 |
| Length | 137 |
| Max. P | 0.574617 |

| Location | 13,424,186 – 13,424,287 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 89.28 |
| Mean single sequence MFE | -31.04 |
| Consensus MFE | -22.32 |
| Energy contribution | -23.30 |
| Covariance contribution | 0.98 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.539807 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13424186 101 - 27905053 AAUUAACCCAUCAGACGGGCUUAUGAGCCGCCGACCGCAAAUGCACGCGCACACACGCAAUUGCAACUAUGCAAUUAUCAAAUUAGUGCGGGGCGUGGCGG ......(((.......)))........((((((..(((...((((((((......)))(((((((....))))))).........)))))..))))))))) ( -34.40) >DroSec_CAF1 49809 101 - 1 AAUUAACCCAUCAGACGGGCUUAUGAGCCGUCGACCGCAAAUGCACACGCACACACGCAAUUGCAACUAUGCAAUUAUCAAAUUAGUGCGGGGCGUGGCGG .............(((((.((....)))))))..((((..((((...(((((......(((((((....))))))).........)))))..)))).)))) ( -32.16) >DroSim_CAF1 52627 101 - 1 AAUUAACCCAUCAGACGGGCUUAUGAGCCGUCGACCGCAAAUGCACACGCACACACGCAAUUGCAACUAUGCAAUUAUCAAAUUAGUGCGGGGCGUGGCGG .............(((((.((....)))))))..((((..((((...(((((......(((((((....))))))).........)))))..)))).)))) ( -32.16) >DroEre_CAF1 50105 89 - 1 AAUUAACCCAUCAGACGGGCUUAUGAGCCGCCAAGCGCUAGUGCG----CACACACGCAGUUGC--------AAUUAUCAAAUUAGUGCGGGGCGUGGCGG ......(((.......)))........(((((..((((....)))----)....((((..((((--------(.(((......)))))))).))))))))) ( -29.80) >DroYak_CAF1 54959 89 - 1 AAUUAACCCAUCAGACGGGCUUAUGAGCCGCCGACCGCCAAUGCA----CACACACGCAAUUGC--------AAUUAUCAAAUUAGUGCGGGGCGUGGCGG ......(((.......)))........((((((..((((..((((----(......((....))--------((((....)))).))))).)))))))))) ( -26.70) >consensus AAUUAACCCAUCAGACGGGCUUAUGAGCCGCCGACCGCAAAUGCAC_CGCACACACGCAAUUGCAACUAUGCAAUUAUCAAAUUAGUGCGGGGCGUGGCGG ......(((.......)))........((((((..(((...(((((..((......))(((((((....))))))).........)))))..))))))))) (-22.32 = -23.30 + 0.98)



| Location | 13,424,225 – 13,424,323 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 89.67 |
| Mean single sequence MFE | -23.04 |
| Consensus MFE | -18.58 |
| Energy contribution | -18.78 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.574617 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
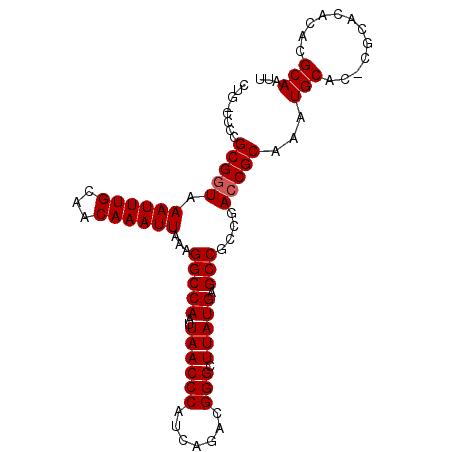
>3R_DroMel_CAF1 13424225 98 - 27905053 CUGGCCGCGCGGUAAAUUUGCAACAAAUUAAAGGCCAAUUAACCCAUCAGACGGGCUUAUGAGCCGCCGACCGCAAAUGCACGCGCACACACGCAAUU ......(((((....((((((...........(((((..((((((.......))).))))).)))(.....)))))))...)))))............ ( -24.50) >DroSec_CAF1 49848 92 - 1 ------CCGCGGUAAAUUUGCAACAAAUUAAAGGCCAAUUAACCCAUCAGACGGGCUUAUGAGCCGUCGACCGCAAAUGCACACGCACACACGCAAUU ------..(((((..((((((...........((........)).....(((((.((....)))))))....))))))..)).)))............ ( -22.20) >DroSim_CAF1 52666 92 - 1 ------CCGCGGUAAAUUUGCAACAAAUUAAAGGCCAAUUAACCCAUCAGACGGGCUUAUGAGCCGUCGACCGCAAAUGCACACGCACACACGCAAUU ------..(((((..((((((...........((........)).....(((((.((....)))))))....))))))..)).)))............ ( -22.20) >DroEre_CAF1 50136 93 - 1 CUG-CCCCGCGGUAAAUUUGCAACAAAUUAAAGGCCAAUUAACCCAUCAGACGGGCUUAUGAGCCGCCAAGCGCUAGUGCG----CACACACGCAGUU (((-(...(((((.((((((...))))))..(((((..((.........))..)))))....)))))...((((....)))----)......)))).. ( -26.00) >DroYak_CAF1 54990 94 - 1 CUGACCCCGCGGUAAAUUUGCAACAAAUUAAAGGCCAAUUAACCCAUCAGACGGGCUUAUGAGCCGCCGACCGCCAAUGCA----CACACACGCAAUU ........(((((.((((((...))))))...(((((..((((((.......))).))))).)))....)))))...(((.----.......)))... ( -20.30) >consensus CUG_CCCCGCGGUAAAUUUGCAACAAAUUAAAGGCCAAUUAACCCAUCAGACGGGCUUAUGAGCCGCCGACCGCAAAUGCAC_CGCACACACGCAAUU ........(((((.((((((...))))))...(((((..((((((.......))).))))).)))....)))))...(((............)))... (-18.58 = -18.78 + 0.20)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:56 2006