| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
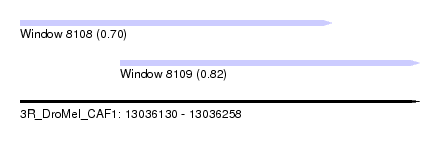
| Location | 13,036,130 – 13,036,258 |
| Length | 128 |
| Max. P | 0.823997 |

| Location | 13,036,130 – 13,036,230 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 78.57 |
| Mean single sequence MFE | -36.72 |
| Consensus MFE | -20.24 |
| Energy contribution | -20.37 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.696983 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13036130 100 + 27905053 GCG-GCCAAACGAAAUGCAUGAGACAGCUUAAGUGAGCUGCGCUGCUGCUACACAAAAGUGUGCGGUGGGCGGUGGGCGGCUGGGC------CAGCAGUUGCGGCAG .((-(((............((((....)))).((..(((((.(..((((.((((....))))))))..))))))..))))))).((------(.((....))))).. ( -40.60) >DroSec_CAF1 56197 93 + 1 GCGGGCCAAACGAAAUGCAUGAGACAGCUUAAGUGAGCUGCGCUGCUGCUUCACAAAAGU-------GGGCGGUGGGCGGUUGGGG------CAGCAG-UGCGGCAG ....(((..................(((((....)))))((((((((((((((((....)-------).((.....))...)))))------))))))-)))))).. ( -35.80) >DroSim_CAF1 48425 92 + 1 GCG-GCCAAACGAAAUGCAUGAGACAGCUUAAGUGAGCUGCGCUGCUGCUUCACAAAAGU-------GGGCGGUGGGCUGCUGGGG------CAGCAG-UGCGGCAG ...-(((..................(((((....)))))(((((((((((((((....))-------.(((((....)))))))))------))))))-)))))).. ( -40.30) >DroEre_CAF1 56400 88 + 1 GCG-GCCAAACGAAAUGCAUGAGACAGCUUAAGUGAGCUGCGCUGCUGCUCCGCAAAAG--------GGGCAGUGGGCGGAGGGC---------GUUG-GACGGUUG .((-.((((.((...(((....(.((((((....))))))).(..(((((((......)--------))))))..)))).....)---------))))-).)).... ( -35.30) >DroAna_CAF1 55691 88 + 1 AAG-GCCAAACGAAAUGCAUGAGACAGCUUAAGUGAGCUGCGC-GCCGCUAC-----CGU-------CGGAGGUG--CGGAUAGGGGCGUCCCAGUAG---UGGCAG ...-((((.((...........(.((((((....)))))))((-(((.((((-----(((-------.......)--))).))).)))))....))..---)))).. ( -31.60) >consensus GCG_GCCAAACGAAAUGCAUGAGACAGCUUAAGUGAGCUGCGCUGCUGCUACACAAAAGU_______GGGCGGUGGGCGGAUGGGG______CAGCAG_UGCGGCAG ....(((........(((......((((((....))))))((((((((((..................)))))).))))...............))).....))).. (-20.24 = -20.37 + 0.13)



| Location | 13,036,162 – 13,036,258 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 82.98 |
| Mean single sequence MFE | -39.08 |
| Consensus MFE | -24.36 |
| Energy contribution | -25.17 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.823997 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 13036162 96 + 27905053 UGAGCUGCGCUGCUGCUACACAAAAGUGUGCGGUGGGCGGUGGGCGGCUGGGCCAGCAGUUGCGGCAGGGGGCGUGGCAGCGGGCAAAAUGCGGAU---- ...(((((.(..((((.((((....))))))))..))))))..(((.....(((.((.((..((.(....).))..)).)).)))....)))....---- ( -41.80) >DroSec_CAF1 56230 88 + 1 UGAGCUGCGCUGCUGCUUCACAAAAGU-------GGGCGGUGGGCGGUUGGGGCAGCAG-UGCGGCAGGGGGCGUGGCAGCGGGCAAAAUGCGGAU---- ...(((((((((((((((((((....)-------).((.....))...)))))))))))-)))))).....((((.((.....))...))))....---- ( -38.70) >DroSim_CAF1 48457 88 + 1 UGAGCUGCGCUGCUGCUUCACAAAAGU-------GGGCGGUGGGCUGCUGGGGCAGCAG-UGCGGCAGGGGGCGUGGCAGCGGGCAAAAUGCGGAU---- ...((((((((((((((((((....))-------.(((((....)))))))))))))))-)))))).....((((.((.....))...))))....---- ( -43.20) >DroEre_CAF1 56432 88 + 1 UGAGCUGCGCUGCUGCUCCGCAAAAG--------GGGCAGUGGGCGGAGGGC---GUUG-GACGGUUGGGGGCGUGGCAGCGGGCGAAAUGCGGAUGUGG ....((((.(..(((((((......)--------))))))..)))))...((---(((.-..(.((((.........)))).)....)))))........ ( -32.60) >consensus UGAGCUGCGCUGCUGCUUCACAAAAGU_______GGGCGGUGGGCGGCUGGGGCAGCAG_UGCGGCAGGGGGCGUGGCAGCGGGCAAAAUGCGGAU____ ...(((.((((((((((((((....)).........((.((...((((((...))))))..)).))..)))))..))))))))))............... (-24.36 = -25.17 + 0.81)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:42:56 2006