
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,645,435 – 12,645,529 |
| Length | 94 |
| Max. P | 0.998360 |

| Location | 12,645,435 – 12,645,529 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 96.07 |
| Mean single sequence MFE | -42.50 |
| Consensus MFE | -39.12 |
| Energy contribution | -38.96 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.92 |
| SVM decision value | 3.08 |
| SVM RNA-class probability | 0.998360 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
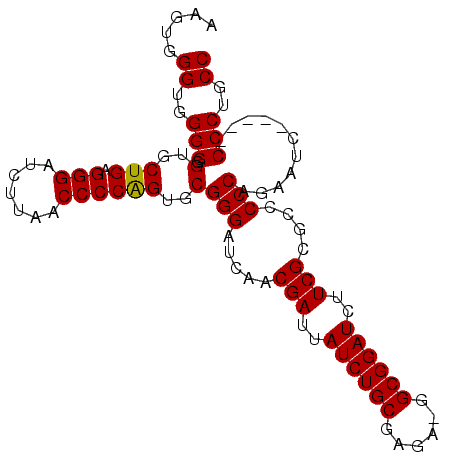
>3R_DroMel_CAF1 12645435 94 + 27905053 AAGUGGGUGGGGGGUGCUGAGGGAUCUUAACCCCAGUGCGGGAUCAACGAUUAUCUGCCAGA-GGCGGAUCUUCGCGCCCCCAGAAUC-----CCCUGCC .....((..(((((..(((.(((.......))))))..)(((.....(((..(((((((...-)))))))..)))....))).....)-----)))..)) ( -43.60) >DroSec_CAF1 21917 94 + 1 AAGUGGGUGGGGGGUGCUGAGGGAUCUUAACCCCAGUGCGGGAUCAACGAUUAUCUGCGAGA-GGCGGAUCUUCGCGCCCCCAGAAUC-----CCCUGCC .....((..(((((..(((.(((.......))))))..)(((.....(((..((((((....-.))))))..)))....))).....)-----)))..)) ( -42.00) >DroSim_CAF1 19880 94 + 1 AAGUGGGUGGGGGGUGCUGAGGGAUCUUAACCCCAGUGCGGGAUCAACGAUUAUCUGCGAGA-GGCGGAUCUUCGCGCCCCCAGAAUC-----CCCUGCC .....((..(((((..(((.(((.......))))))..)(((.....(((..((((((....-.))))))..)))....))).....)-----)))..)) ( -42.00) >DroEre_CAF1 25203 95 + 1 AAGUGGGUGGGGGGUGCUGAGGGAUCUUAACCCCAGUGCGGGAUCAACGAUUAUCUGCGAGAUGGCGGAUCUUCGCGCCCCCAGAAUC-----CCCUGCC .....((..(((((..(((.(((.......))))))..)(((.....(((..((((((......))))))..)))....))).....)-----)))..)) ( -42.20) >DroYak_CAF1 13714 100 + 1 AAGUGGGUGGGGGGUGCUGAGGGAUCUUAACCCCGGUGCGGGAUCAACGAUUAUCUGCGAGAUGGCGGAUCUUCGCGCCCCCAGAAUGACGCGCCCUGCC ....(((((((((((((.((((((((((.((....))..)))))).......((((((......)))))))))))))))))).(.....).))))).... ( -42.70) >consensus AAGUGGGUGGGGGGUGCUGAGGGAUCUUAACCCCAGUGCGGGAUCAACGAUUAUCUGCGAGA_GGCGGAUCUUCGCGCCCCCAGAAUC_____CCCUGCC .....((..(((.(..(((.(((.......))))))..)(((.....(((..((((((......))))))..)))....)))...........)))..)) (-39.12 = -38.96 + -0.16)


| Location | 12,645,435 – 12,645,529 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 96.07 |
| Mean single sequence MFE | -37.08 |
| Consensus MFE | -31.55 |
| Energy contribution | -31.55 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.917756 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
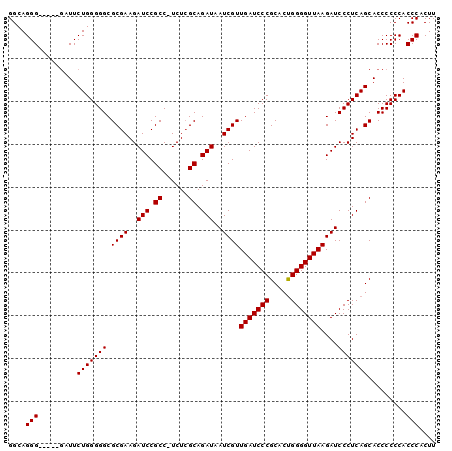
>3R_DroMel_CAF1 12645435 94 - 27905053 GGCAGGG-----GAUUCUGGGGGCGCGAAGAUCCGCC-UCUGGCAGAUAAUCGUUGAUCCCGCACUGGGGUUAAGAUCCCUCAGCACCCCCCACCCACUU ((..(((-----(...(((((((.((((..(((.(((-...))).)))..))))(((((((.....)))))))....)))))))...))))..))..... ( -38.70) >DroSec_CAF1 21917 94 - 1 GGCAGGG-----GAUUCUGGGGGCGCGAAGAUCCGCC-UCUCGCAGAUAAUCGUUGAUCCCGCACUGGGGUUAAGAUCCCUCAGCACCCCCCACCCACUU ((..(((-----(...(((((((.((((.((......-)))))).....(((.((((((((.....))))))))))))))))))...))))..))..... ( -37.00) >DroSim_CAF1 19880 94 - 1 GGCAGGG-----GAUUCUGGGGGCGCGAAGAUCCGCC-UCUCGCAGAUAAUCGUUGAUCCCGCACUGGGGUUAAGAUCCCUCAGCACCCCCCACCCACUU ((..(((-----(...(((((((.((((.((......-)))))).....(((.((((((((.....))))))))))))))))))...))))..))..... ( -37.00) >DroEre_CAF1 25203 95 - 1 GGCAGGG-----GAUUCUGGGGGCGCGAAGAUCCGCCAUCUCGCAGAUAAUCGUUGAUCCCGCACUGGGGUUAAGAUCCCUCAGCACCCCCCACCCACUU ((..(((-----(...(((((((.((((.(((.....))))))).....(((.((((((((.....))))))))))))))))))...))))..))..... ( -37.70) >DroYak_CAF1 13714 100 - 1 GGCAGGGCGCGUCAUUCUGGGGGCGCGAAGAUCCGCCAUCUCGCAGAUAAUCGUUGAUCCCGCACCGGGGUUAAGAUCCCUCAGCACCCCCCACCCACUU ((..(((.(.((....(((((((.((((.(((.....))))))).....(((.(((((((((...))))))))))))))))))).)).))))..)).... ( -35.00) >consensus GGCAGGG_____GAUUCUGGGGGCGCGAAGAUCCGCC_UCUCGCAGAUAAUCGUUGAUCCCGCACUGGGGUUAAGAUCCCUCAGCACCCCCCACCCACUU ....(((.........(((((((.((((..(((.((......)).)))..))))(((((((.....)))))))....))))))).........))).... (-31.55 = -31.55 + 0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:59 2006