
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,642,228 – 12,642,366 |
| Length | 138 |
| Max. P | 0.761527 |

| Location | 12,642,228 – 12,642,338 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.65 |
| Mean single sequence MFE | -38.32 |
| Consensus MFE | -24.01 |
| Energy contribution | -24.57 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.65 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.761527 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12642228 110 - 27905053 ----GGAGUCUC-UUUGGCGCCACUUCAUUGUGAUCAUUGAAUGGCCGUUGAUUGUGUCGCAUCGAUUCGCGGCGAUUUUAAUCGCAGGCGGCGGACAUUAA-----ACGCCUUUAAUUU ----((.(((..-...))).))......(((((((....((((.(((((.(((((........))))).))))).))))..)))))))..((((........-----.))))........ ( -34.30) >DroPse_CAF1 16233 115 - 1 ----AGAGUCGC-UCUGGCGCCACUUCAUUGUGAUUAUCGAAUGGCCGUUGAUUGUGUGGCAUCGAUCCGCGGCGAUUUGAAUCGCAGCCGGAGCACAUUAAAAUGAACGCCUUUCAUUU ----...((.((-(((((((((((......)))....((((((.(((((.(((((........))))).))))).))))))...)).))))))))))....(((((((.....))))))) ( -45.30) >DroGri_CAF1 42091 113 - 1 ACACACAGACAUAUUGGGCACCACUCCAUUGUCAUCAUUGAGAGGCCGUUGAUUG--UCGUGUCGAUUCGCGGCGAUUUUAAUCGCAACUGGUGCGCAUUAA-----ACGCCUUUCAUUU .......((((...((((......)))).)))).....((((((((.((((((((--(((((......)))))))))).((((((((.....)))).)))))-----))))))))))... ( -37.00) >DroWil_CAF1 35288 106 - 1 ------ACUCAC-AAUGGCGCCACUUCAUUGUCAUCAUGGAAUUGCCGUUGAUUG--UCGCAUCGAUUCGCGGCGAUUUUAAUCGCAAUGGGCAAGCAUUAA-----ACGCCUUUCAUUU ------......-...((((((....((((((.((..((((((((((((.(((((--......))))).)))))))))))))).)))))))))..(......-----.))))........ ( -30.40) >DroAna_CAF1 9079 110 - 1 ----AGAGUCGC-UGUGGCGCCACUUCAUUGUGAUCAUUGAAUGGCCGUUGAUUGUGUCGCAUCGAUUCGCGGCGAUUUUAAUCGCAAGCGGCGCACAUUAA-----ACGCCUUUCAUUU ----.(((..((-.((((((((.((....((((((....((((.(((((.(((((........))))).))))).))))..)))))))).))))).)))...-----..))..))).... ( -37.60) >DroPer_CAF1 20923 115 - 1 ----AGAGUCGC-UCUGGCGCCACUUCAUUGUGAUUAUCGAAUGGCCGUUGAUUGUGUCGCAUCGAUCCGCGGCGAUUUGAAUCGCAGCCGGAGCACAUUAAAAUGAACGCCUUUCAUUU ----...((.((-(((((((((((......)))....((((((.(((((.(((((........))))).))))).))))))...)).))))))))))....(((((((.....))))))) ( -45.30) >consensus ____AGAGUCGC_UCUGGCGCCACUUCAUUGUGAUCAUUGAAUGGCCGUUGAUUGUGUCGCAUCGAUUCGCGGCGAUUUUAAUCGCAACCGGCGCACAUUAA_____ACGCCUUUCAUUU .................(((((......(((((((....((((.(((((.(((((........))))).))))).))))..)))))))..)))))......................... (-24.01 = -24.57 + 0.56)



| Location | 12,642,263 – 12,642,366 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 76.15 |
| Mean single sequence MFE | -27.20 |
| Consensus MFE | -15.45 |
| Energy contribution | -15.90 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.577277 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12642263 103 - 27905053 GGACAACUAAUUCCACUCACAU----ACAGAUGGAGUCUCUUUGGCGCCACUUCAUUGUGAUCAUUGAAUGGCCGUUGAUUGUGUCGCAUCGAUUCGCGGCGAUUUU (((........)))..(((((.----..((.(((.(((.....))).)))))....))))).....((((.(((((.(((((........))))).))))).)))). ( -29.00) >DroPse_CAF1 16273 87 - 1 ---C-------UACCCUCACA----------AAGAGUCGCUCUGGCGCCACUUCAUUGUGAUUAUCGAAUGGCCGUUGAUUGUGUGGCAUCGAUCCGCGGCGAUUUG ---.-------.....(((((----------(.(((((((....)))..))))..))))))....(((((.(((((.(((((........))))).))))).))))) ( -29.00) >DroEre_CAF1 21997 107 - 1 GCACAACUAAUUCCACUCACAUAUGUACAGAUGGAGUCUCUUUGGCGCCACUUCAUUGUGAUCAUUGAAUGCCAGUUGAUUGUGUCGCAUCGAUUCGCGGCGAUUUU ...(((((.((((...(((((.(((...((.(((.(((.....))).))))).)))))))).....))))...)))))....((((((........))))))..... ( -26.70) >DroWil_CAF1 35323 81 - 1 ------------ACACUCACAA------------ACUCACAAUGGCGCCACUUCAUUGUCAUCAUGGAAUUGCCGUUGAUUG--UCGCAUCGAUUCGCGGCGAUUUU ------------..........------------....(((((((.......)))))))......(((((((((((.(((((--......))))).))))))))))) ( -21.40) >DroAna_CAF1 9114 91 - 1 AGACAACCAAUUCCA----------------UAGAGUCGCUGUGGCGCCACUUCAUUGUGAUCAUUGAAUGGCCGUUGAUUGUGUCGCAUCGAUUCGCGGCGAUUUU ...............----------------.(((((((((((((.((((.((((.((....)).)))))))).((((((.(.....)))))))))))))))))))) ( -28.10) >DroPer_CAF1 20963 86 - 1 -----------UACCCUCACA----------AAGAGUCGCUCUGGCGCCACUUCAUUGUGAUUAUCGAAUGGCCGUUGAUUGUGUCGCAUCGAUCCGCGGCGAUUUG -----------.....(((((----------(.(((((((....)))..))))..))))))....(((((.(((((.(((((........))))).))))).))))) ( -29.00) >consensus ___C_______UACACUCACA__________UAGAGUCGCUCUGGCGCCACUUCAUUGUGAUCAUUGAAUGGCCGUUGAUUGUGUCGCAUCGAUUCGCGGCGAUUUU ...................................(((.....)))..(((......)))......((((.(((((.(((((........))))).))))).)))). (-15.45 = -15.90 + 0.45)


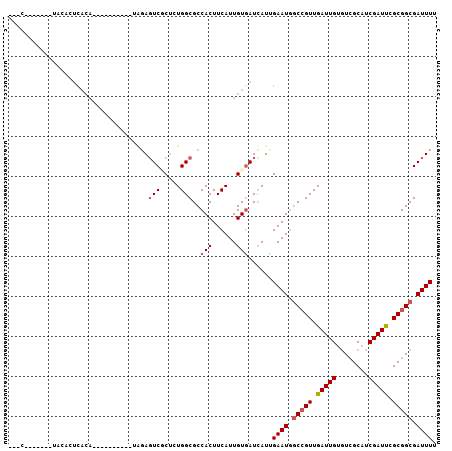
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:57 2006