
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,527,141 – 12,527,250 |
| Length | 109 |
| Max. P | 0.985274 |

| Location | 12,527,141 – 12,527,250 |
|---|---|
| Length | 109 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 76.81 |
| Mean single sequence MFE | -30.94 |
| Consensus MFE | -15.46 |
| Energy contribution | -16.68 |
| Covariance contribution | 1.22 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.50 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.822084 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
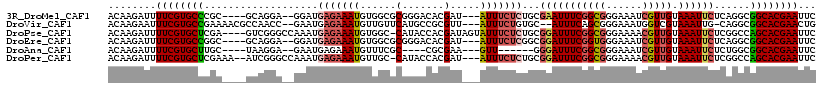
>3R_DroMel_CAF1 12527141 109 + 27905053 GAAUUCGUGCCGCCUGAGAAUUUACAACGAUUUCCCGCCGAAAUUCGCAGAGAAAU---AUCGUGUCCCGCGCCACAUUUCUCAUCC--UCCUGC----GCGGGCACGAAAAUCUUGU ...(((((((((((.(((.........(((((((.....)))).)))..(((((((---...(((.....)))...)))))))...)--))..).----)).))))))))........ ( -30.80) >DroVir_CAF1 8626 110 + 1 CAGUUCGUGCCGCCUG-CAAUUUACGACCAUUUCCCGCUGAAAU--GCACAGAAAU---AACGCGGCAUGAACAACAUUUCUCAUUC--GGUUGGCGUUUUCGGCACGAAAUUCUUGU ..(((((((((((.((-((.((((((.........)).)))).)--))).(....)---...)))))))))))...........(((--((((((.....))))).))))........ ( -30.70) >DroPse_CAF1 15469 113 + 1 GAAUUCGUGCUGGCCGAGAAUUUACAACGUUUUCCCGCCGAAAUCCGCAGAGAAAUACUAUCGUGGUAUG-GCCACAUUUCUCAUUUGGCCCGAC----UCGAGCACGAAAAUCUUGU ...(((((((((((((((..................((.(....).)).(((((((......((((....-.))))))))))).)))))))....----...))))))))........ ( -32.81) >DroEre_CAF1 13793 109 + 1 GAAUUCGUGCCGCCUGAGAAUUUACAACGAUUUCCCACCGAAAUCCGCCGAGAAAU---AUCGUGUCCCGCGCCACAUUUCUCAUCC--UCCUGC----GCCGGCACGAAAAUCUUGU ...(((((((((((.(((..........((((((.....))))))....(((((((---...(((.....)))...)))))))...)--))..).----).)))))))))........ ( -32.50) >DroAna_CAF1 16006 99 + 1 GAAUUCGUGCCGCCAGAGAAUUUACAACGAUUUCCCGCCGAAAUCCC------AAC---UUCGCG----GCGAAACAUUUCUCAUUC--UCCUUA----GCAAGCACGAAAAUCUUGU ...(((((((.((..((((((.........((((((((.(((.....------...---))))))----).))))........))))--))....----))..)))))))........ ( -25.63) >DroPer_CAF1 19434 112 + 1 GAAUUCGUGCUGGCCGAGAAUUUACAACGUUUUCCCGCCGAAAUCCGCAGAGAAAU---AUCGUGGUAUG-GCAACAUUUCUCAUUUGGCCCGAU--UUUCGAGCACGAAAAUCUUGU ...((((((((((((((((((.......))))))..((((....((((.((.....---.))))))..))-))..............))))((..--...))))))))))........ ( -33.20) >consensus GAAUUCGUGCCGCCUGAGAAUUUACAACGAUUUCCCGCCGAAAUCCGCAGAGAAAU___AUCGUGGCAUGCGCAACAUUUCUCAUUC__UCCUGC____GCGAGCACGAAAAUCUUGU ...((((((((.................((((((.....))))))....(((((((.....((.....))......)))))))...................))))))))........ (-15.46 = -16.68 + 1.22)



| Location | 12,527,141 – 12,527,250 |
|---|---|
| Length | 109 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 76.81 |
| Mean single sequence MFE | -38.25 |
| Consensus MFE | -20.09 |
| Energy contribution | -20.73 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.53 |
| SVM decision value | 2.00 |
| SVM RNA-class probability | 0.985274 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12527141 109 - 27905053 ACAAGAUUUUCGUGCCCGC----GCAGGA--GGAUGAGAAAUGUGGCGCGGGACACGAU---AUUUCUCUGCGAAUUUCGGCGGGAAAUCGUUGUAAAUUCUCAGGCGGCACGAAUUC ...(((...((((((((((----((....--(.((.....)).).))))))).))))).---...)))(((.((((((((((((....)))))).)))))).)))............. ( -38.10) >DroVir_CAF1 8626 110 - 1 ACAAGAAUUUCGUGCCGAAAACGCCAACC--GAAUGAGAAAUGUUGUUCAUGCCGCGUU---AUUUCUGUGC--AUUUCAGCGGGAAAUGGUCGUAAAUUG-CAGGCGGCACGAACUG ........(((((((((....)(((....--(((..(......)..))).(((.(((((---((((((.(((--......))))))))))).))).....)-)))))))))))))... ( -31.90) >DroPse_CAF1 15469 113 - 1 ACAAGAUUUUCGUGCUCGA----GUCGGGCCAAAUGAGAAAUGUGGC-CAUACCACGAUAGUAUUUCUCUGCGGAUUUCGGCGGGAAAACGUUGUAAAUUCUCGGCCAGCACGAAUUC ........((((((((.(.----(((((((((.((.....)).))))-).....(((((.((.((((.((((.(....).))))))))))))))).......)))))))))))))... ( -38.00) >DroEre_CAF1 13793 109 - 1 ACAAGAUUUUCGUGCCGGC----GCAGGA--GGAUGAGAAAUGUGGCGCGGGACACGAU---AUUUCUCGGCGGAUUUCGGUGGGAAAUCGUUGUAAAUUCUCAGGCGGCACGAAUUC ........(((((((((.(----....((--(((((((((((((.(.(.....).).))---)))))))((((.(((((.....)))))))))....)))))).).)))))))))... ( -41.30) >DroAna_CAF1 16006 99 - 1 ACAAGAUUUUCGUGCUUGC----UAAGGA--GAAUGAGAAAUGUUUCGC----CGCGAA---GUU------GGGAUUUCGGCGGGAAAUCGUUGUAAAUUCUCUGGCGGCACGAAUUC ........((((((((.((----(..(((--((((....((((((((.(----((((((---((.------...))))).)))))))).))))....))))))))))))))))))... ( -38.30) >DroPer_CAF1 19434 112 - 1 ACAAGAUUUUCGUGCUCGAAA--AUCGGGCCAAAUGAGAAAUGUUGC-CAUACCACGAU---AUUUCUCUGCGGAUUUCGGCGGGAAAACGUUGUAAAUUCUCGGCCAGCACGAAUUC ........((((((((((...--..))((((....(((((((((((.-.......))))---))))))).(.(((((((((((......))))).)))))).)))))))))))))... ( -41.90) >consensus ACAAGAUUUUCGUGCCCGA____GCAGGA__GAAUGAGAAAUGUGGCGCAUGCCACGAU___AUUUCUCUGCGGAUUUCGGCGGGAAAUCGUUGUAAAUUCUCAGGCGGCACGAAUUC ........((((((((...................(((((((......(.......).....)))))))...(((((((((((......))))).))))))......))))))))... (-20.09 = -20.73 + 0.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:06 2006