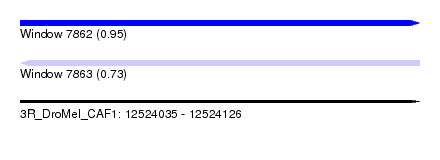

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,524,035 – 12,524,126 |
| Length | 91 |
| Max. P | 0.951529 |

| Location | 12,524,035 – 12,524,126 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 80.35 |
| Mean single sequence MFE | -32.26 |
| Consensus MFE | -22.70 |
| Energy contribution | -22.95 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.951529 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12524035 91 + 27905053 ---------C-GCUGCAGUUG------CUGCUGCGUUAAUGGCCUGGGUAAUUGCUUUUAAUUGCCAUUGUAAAGUUUUUCGCAUCGCAACCAAAAAUGGCAGCGGC------------ ---------(-(((((.((((------(.(.((((..(((.((...((((((((....))))))))...))...)))...)))).))))))((....))))))))..------------ ( -30.90) >DroPse_CAF1 12169 98 + 1 ---------CUGCUGCCUGUGCCGCCGCUGCUGCGUUAAUGACUUUGGUAAUUACUUUUAAUUGCCAUUGUAAAGCUUUUCGCAUCGCCACCAAAAAUGGUAGCGGC------------ ---------..((.......)).((((((((((((......((..(((((((((....)))))))))..)).........))))..)).((((....))))))))))------------ ( -29.06) >DroYak_CAF1 9899 91 + 1 ---------C-GCUGCGGUUG------CCGCUGCGUUAAUGGCCUGGGUAAUUGCUUUUAAUUGCCAUUGUAAAGUUUUUCGCAUCGCAACCGAAAAUGGCAGCGGC------------ ---------(-((((((((((------(.(.((((..(((.((...((((((((....))))))))...))...)))...)))).))))))).......))))))..------------ ( -33.11) >DroMoj_CAF1 4986 110 + 1 UUUUGUUGUC-GCUGCUGCUG------CUGCUGCGUUAAUGGCUGUGGUAAUUACUUUUAAUUGCCAUUGUAAAGUU-UUCGCAUCGC-ACCAAAAAUGGCAGCGCCUCAGCAUCUUGG ...(((((.(-(((((((.((------.(((((((..(((.((.((((((((((....)))))))))).))...)))-..))))..))-).))....))))))))...)))))...... ( -37.80) >DroAna_CAF1 13351 98 + 1 --UGACUGCC-ACUGCUGCUG------CUGCUGCGUUAAUGGCUUUGGUAAUUACUUUUAAUUGCCAUUGUAAAGUUUUUCGCAUCGCAACCGAAAAUGGCAGCAGC------------ --........-...(((((((------((..((((..(((.((..(((((((((....)))))))))..))...)))...))))(((....)))....)))))))))------------ ( -34.00) >DroPer_CAF1 16061 92 + 1 ---------CUGCUGCCUGUG------CUGCUGCGUUAAUGACUCUGGUAAUUACUUUUAAUUGCCAUUGUAAAGCUUUUCGCAUCGCCACCAAAAAUGGUAGCGGC------------ ---------((((((((.(((------.((.((((......((..(((((((((....)))))))))..)).........)))).)).))).......)))))))).------------ ( -28.66) >consensus _________C_GCUGCCGCUG______CUGCUGCGUUAAUGGCUUUGGUAAUUACUUUUAAUUGCCAUUGUAAAGUUUUUCGCAUCGCAACCAAAAAUGGCAGCGGC____________ ...........((((((...........((.((((..(((.((..(((((((((....)))))))))..))...)))...)))).))...........))))))............... (-22.70 = -22.95 + 0.25)
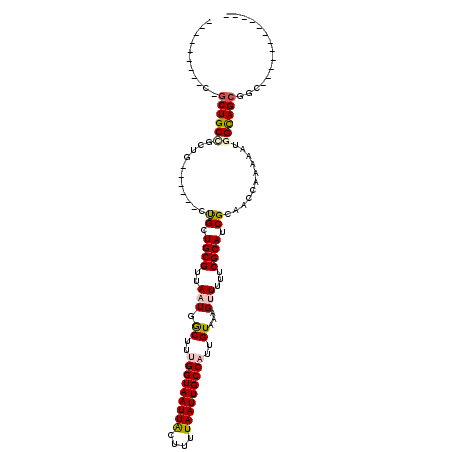


| Location | 12,524,035 – 12,524,126 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 80.35 |
| Mean single sequence MFE | -27.72 |
| Consensus MFE | -15.86 |
| Energy contribution | -15.45 |
| Covariance contribution | -0.41 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.729323 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12524035 91 - 27905053 ------------GCCGCUGCCAUUUUUGGUUGCGAUGCGAAAAACUUUACAAUGGCAAUUAAAAGCAAUUACCCAGGCCAUUAACGCAGCAG------CAACUGCAGC-G--------- ------------..((((((.......((((((..((((((.....))..((((((......((....))......))))))..))))...)------))))))))))-)--------- ( -26.51) >DroPse_CAF1 12169 98 - 1 ------------GCCGCUACCAUUUUUGGUGGCGAUGCGAAAAGCUUUACAAUGGCAAUUAAAAGUAAUUACCAAAGUCAUUAACGCAGCAGCGGCGGCACAGGCAGCAG--------- ------------((((((((((....)))))))(((((.....)).......(((.(((((....))))).)))..)))......((.((....)).))...))).....--------- ( -27.80) >DroYak_CAF1 9899 91 - 1 ------------GCCGCUGCCAUUUUCGGUUGCGAUGCGAAAAACUUUACAAUGGCAAUUAAAAGCAAUUACCCAGGCCAUUAACGCAGCGG------CAACCGCAGC-G--------- ------------((((((((..((((((.........)))))).......((((((......((....))......))))))...)))))))------).........-.--------- ( -28.30) >DroMoj_CAF1 4986 110 - 1 CCAAGAUGCUGAGGCGCUGCCAUUUUUGGU-GCGAUGCGAA-AACUUUACAAUGGCAAUUAAAAGUAAUUACCACAGCCAUUAACGCAGCAG------CAGCAGCAGC-GACAACAAAA ....(.(((((..(((((((..((..((((-((...))...-..........(((.(((((....))))).)))..))))..)).))))).)------)..))))).)-.......... ( -28.00) >DroAna_CAF1 13351 98 - 1 ------------GCUGCUGCCAUUUUCGGUUGCGAUGCGAAAAACUUUACAAUGGCAAUUAAAAGUAAUUACCAAAGCCAUUAACGCAGCAG------CAGCAGCAGU-GGCAGUCA-- ------------...(((((((((....(((((..((((((.....))..((((((......((....))......))))))..))))...)------))))...)))-))))))..-- ( -29.70) >DroPer_CAF1 16061 92 - 1 ------------GCCGCUACCAUUUUUGGUGGCGAUGCGAAAAGCUUUACAAUGGCAAUUAAAAGUAAUUACCAGAGUCAUUAACGCAGCAG------CACAGGCAGCAG--------- ------------((((((((((....)))))))(.((((....((((.....(((.(((((....))))).)))))))......)))).)..------....))).....--------- ( -26.00) >consensus ____________GCCGCUGCCAUUUUUGGUUGCGAUGCGAAAAACUUUACAAUGGCAAUUAAAAGUAAUUACCAAAGCCAUUAACGCAGCAG______CAACAGCAGC_G_________ ............(((((.(((......))).))).((((((.....))..((((((......((....))......))))))..))))))............................. (-15.86 = -15.45 + -0.41)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:02 2006