
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,513,773 – 12,513,935 |
| Length | 162 |
| Max. P | 0.998523 |

| Location | 12,513,773 – 12,513,865 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 72.78 |
| Mean single sequence MFE | -35.50 |
| Consensus MFE | -12.98 |
| Energy contribution | -12.18 |
| Covariance contribution | -0.80 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.37 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.923286 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12513773 92 + 27905053 CGCAGCC-UG------AAAACUCGAAAUGAAAUUAAACCCUUUUCGGCGGCUGCUGCUG---CCGCUGACGCUGUUGCAG------CAACAUGGCAACAUGGCGGCUG .((((((-.(------(....))(...(((((.........))))).)))))))....(---((((....((((...)))------)..((((....))))))))).. ( -33.40) >DroPse_CAF1 1005 103 + 1 CGCAACG-UGCAACGCGAAACUCGAAAUGAAAUUAAACCCUUUUUGGCGGCGGUGGC---GGCUGCUGCUGCUUCUGCAAAAGACGGACCGC-GGGACGAGGCAAAUA .((..((-(.(..((((....((.....)).......((.(((((....((((.(((---(((....)))))).)))).))))).))..)))-)).)))..))..... ( -31.80) >DroSec_CAF1 2866 92 + 1 CGCAGCCUUG------AAAACUCGAAAUGAAAUUAAACCCUUUUUCGGCGCUGCUGCUG----CGCUGCCGCUAUUGCAG------CAACAUGGCAACAUGGCGGCUG .(((((.(((------((((.....................))))))).))))).((((----((((((.......))))------)..((((....))))))))).. ( -34.20) >DroSim_CAF1 2930 92 + 1 CGCAGCC-UG------AAAACUCGAAAUGAAAUUAAACCCUUUUCGGCGGCUGCUGCUG---CCGCUGCCGCUGUUGCAG------CAACAUGGCAACAUGGCGGCUG .((((((-.(------(....))(...(((((.........))))).))))))).((((---(((((((.......))))------)...(((....))))))))).. ( -35.60) >DroPer_CAF1 4867 106 + 1 CGCAACG-UGCAACGCGAAACUCGAAAUGAAAUUAAACCCUUUUUGGCGGCGGUGGCGGCGGCUGCUGCUGCUUCUGCCAAAGACGGACCGC-GGGACGAGGCAAAUA .((..((-(.(..((((....((.....)).......((.((((((((((.((..(((((....)))))..)).)))))))))).))..)))-)).)))..))..... ( -42.50) >consensus CGCAGCC_UG______AAAACUCGAAAUGAAAUUAAACCCUUUUUGGCGGCUGCUGCUG___CCGCUGCCGCUGUUGCAG______CAACAUGGCAACAUGGCGGCUG .(((((...............((((((..............))))))..((....)).......)))))(((((((((...............)))..)))))).... (-12.98 = -12.18 + -0.80)



| Location | 12,513,805 – 12,513,896 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 72.11 |
| Mean single sequence MFE | -32.40 |
| Consensus MFE | -17.68 |
| Energy contribution | -18.44 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.55 |
| SVM decision value | 2.13 |
| SVM RNA-class probability | 0.988690 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12513805 91 + 27905053 CUUUUCGGCGGCUGCUGCUG---CCGCUGACGCUGUUGCAG------CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCU ......(.((((((((((((---(.((.......)).))))------)).((((....))))))))))).)(((((.............)))))...... ( -34.62) >DroPse_CAF1 1043 88 + 1 CUUUUUGGCGGCGGUGGC---GGCUGCUGCUGCUUCUGCAAAAGACGGACCGC-GGGACGAGGCAAAUAGUUGCAAAAAAUUAACAUUAUUG-------- ((.((((.(.((((.(((---(((....)))))).))))......((..(...-.)..)).).)))).))......................-------- ( -24.70) >DroSec_CAF1 2899 90 + 1 CUUUUUCGGCGCUGCUGCUG----CGCUGCCGCUAUUGCAG------CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCU ..........((.(((((((----((((((.......))))------)..((((....))))))))).))).)).(((((((..((....)).))))))) ( -32.40) >DroSim_CAF1 2962 91 + 1 CUUUUCGGCGGCUGCUGCUG---CCGCUGCCGCUGUUGCAG------CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCU ......((((((....))))---))((.(((((....))((------(..((((....))))...)))))).)).(((((((..((....)).))))))) ( -35.70) >DroPer_CAF1 4905 98 + 1 CUUUUUGGCGGCGGUGGCGGCGGCUGCUGCUGCUUCUGCCAAAGACGGACCGC-GGGACGAGGCAAAUAGUUGCAAAAAAUUAACAUUAUUGCAGUUU-U .((((((((((.((..(((((....)))))..)).))))))))))((..(...-.)..))..((((...((((........))))....)))).....-. ( -34.60) >consensus CUUUUUGGCGGCUGCUGCUG___CCGCUGCCGCUGUUGCAG______CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCU .....(((((((.((.(......).)).)))))))((((((.........((((....))))((((....))))...............))))))..... (-17.68 = -18.44 + 0.76)

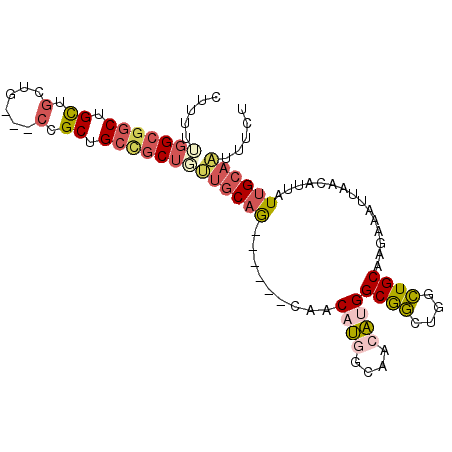

| Location | 12,513,839 – 12,513,935 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 80.57 |
| Mean single sequence MFE | -25.72 |
| Consensus MFE | -18.64 |
| Energy contribution | -18.44 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.19 |
| Mean z-score | -3.32 |
| Structure conservation index | 0.72 |
| SVM decision value | 3.13 |
| SVM RNA-class probability | 0.998523 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12513839 96 + 27905053 GCAG------CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCUGCAGCUUUAACUAACAAGCGCAGAAAAUACACAAAAAAA .(((------(..((((....))))...))))..(((((.............))))).(((((((.((((........)))))))))))............. ( -28.72) >DroSec_CAF1 2932 95 + 1 GCAG------CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCUGCAGCUUUAACUAACAAGCCCAGAGAAUACACAAU-AAA .(((------(..((((....))))...))))((((((.((((((..((....)).))))))))))))..............................-... ( -26.90) >DroSim_CAF1 2996 95 + 1 GCAG------CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCUGCAGCUUUAACUAACAAGCCCAGAGAAUACACAAA-AAA .(((------(..((((....))))...))))((((((.((((((..((....)).))))))))))))..............................-... ( -26.90) >DroEre_CAF1 421 95 + 1 GCAG------CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCUGCAGCUUUAACUAACAAGCCAAGGGAAUACACGAA-AAA .(((------(..((((....))))...))))((((((.((((((..((....)).))))))))))))..............................-... ( -26.90) >DroPer_CAF1 4942 87 + 1 GCCAAAGACGGACCGC-GGGACGAGGCAAAUAGUUGCAAAAAAUUAACAUUAUUGCAGUUU-UGCAGCUUCAACUAGCAAGCCA------UGCCC------- .((......)).....-(((.((.(((.......(((((((...(((.....)))...)))-))))(((......)))..))).------)))))------- ( -19.20) >consensus GCAG______CAACAUGGCAACAUGGCGGCUGGCUGCAAGAAAUUAACAUUAUUGCAAUUUCUGCAGCUUUAACUAACAAGCCCAGAGAAUACACAAA_AAA ((...........((((....))))......(((((((.((((((..((....)).)))))))))))))...........)).................... (-18.64 = -18.44 + -0.20)

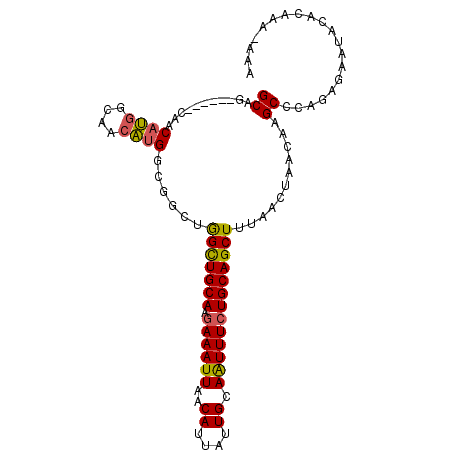
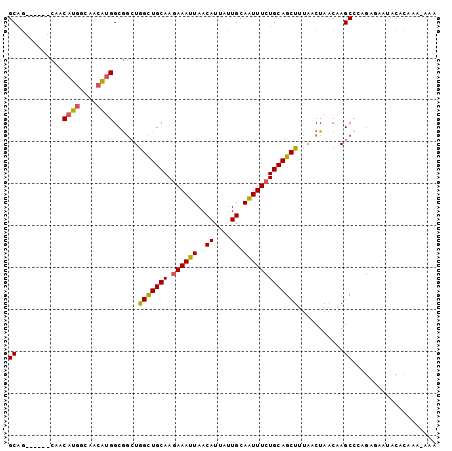
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:38:47 2006