
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,495,958 – 12,496,077 |
| Length | 119 |
| Max. P | 0.999819 |

| Location | 12,495,958 – 12,496,048 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 96.89 |
| Mean single sequence MFE | -24.80 |
| Consensus MFE | -23.14 |
| Energy contribution | -22.82 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.68 |
| SVM RNA-class probability | 0.996287 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12495958 90 - 27905053 UCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGCGUACAACAAUUGAAACGCCAA ..((((((.((.((....(((...(((((....))))).)))...))....))))))))........((((............))))... ( -24.10) >DroSec_CAF1 17452 90 - 1 UCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGCGUACAGCAAUUGAAACGCCAA ..((((((.((.((....(((...(((((....))))).)))...))....))))))))........((((............))))... ( -24.10) >DroSim_CAF1 16766 90 - 1 UCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGCGUACAACAAUUGAAACGCCAA ..((((((.((.((....(((...(((((....))))).)))...))....))))))))........((((............))))... ( -24.10) >DroEre_CAF1 17801 90 - 1 CUGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAACCAAAAGGAAAAGCGCUGGCAGAGGAAAGCGUACAACAAUUGAAACUCCAA ((((((((((..(....)..))..(((((((.((.....))...)))))))..)))))))).(((...((.........))....))).. ( -28.00) >DroYak_CAF1 17730 90 - 1 UUGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGCGUACAACAAUUGAAACUCCAA ((((((((.((.((....(((...(((((....))))).)))...))....)))))))))).(((...((.........))....))).. ( -23.70) >consensus UCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGCGUACAACAAUUGAAACGCCAA ......(((((((...(((((...(((((((..(......)...)))))))))))).((........))))))))).............. (-23.14 = -22.82 + -0.32)



| Location | 12,495,979 – 12,496,077 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 96.94 |
| Mean single sequence MFE | -23.42 |
| Consensus MFE | -20.26 |
| Energy contribution | -20.42 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.957559 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12495979 98 + 27905053 GCUUUCCUCUGCCAGUGCUUUUCCUUUUGCUUUUCCCCAGGAAAACAUGCACUUCGGCACAACUGGCGAAAGAAACCGCAGCUUAUGCUACCCCUCUU ((((((...(((((((..(((((((.............)))))))..(((......)))..)))))))...))))..))(((....)))......... ( -21.82) >DroSec_CAF1 17473 98 + 1 GCUUUCCUCUGCCAGUGCUUUUCCUUUUGCUUUUCCCCAGGAAAACAUGCACUUCGGCACAACUGGCGAAAGAAACCGCAGCUUAUGCUACCCCUCUG ((((((...(((((((..(((((((.............)))))))..(((......)))..)))))))...))))..))(((....)))......... ( -21.82) >DroSim_CAF1 16787 98 + 1 GCUUUCCUCUGCCAGUGCUUUUCCUUUUGCUUUUCCCCAGGAAAACAUGCACUUCGGCACAACUGGCGAAAGAAACCGCAGCUUAUGGUACCCCUCUU (((......((((((((((((((((.............)))))))...)))))..))))......(((........))))))................ ( -20.92) >DroEre_CAF1 17822 98 + 1 GCUUUCCUCUGCCAGCGCUUUUCCUUUUGGUUUUCCCCAGGAAAACAUGCACUUCGGCACAACUGGCAGAAGAAACCGCAGCUUAUGCUACCCCUCGU ((((((.((((((((.(.(((((((...((....))..)))))))).(((......)))...)))))))).))))..))(((....)))......... ( -30.00) >DroYak_CAF1 17751 98 + 1 GCUUUCCUCUGCCAGUGCUUUUCCUUUUGCUUUUCCCCAGGAAAACAUGCACUUCGGCACAACUGGCAAAAGAAACCGCAGCUUAUGCUACCCCUCUU ((((((...(((((((..(((((((.............)))))))..(((......)))..)))))))...))))..))(((....)))......... ( -22.52) >consensus GCUUUCCUCUGCCAGUGCUUUUCCUUUUGCUUUUCCCCAGGAAAACAUGCACUUCGGCACAACUGGCGAAAGAAACCGCAGCUUAUGCUACCCCUCUU ((((((...((((((((.(((((((.............)))))))).(((......)))..)))))))...))))..))(((....)))......... (-20.26 = -20.42 + 0.16)



| Location | 12,495,979 – 12,496,077 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 96.94 |
| Mean single sequence MFE | -35.96 |
| Consensus MFE | -32.54 |
| Energy contribution | -32.62 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.63 |
| Structure conservation index | 0.90 |
| SVM decision value | 4.16 |
| SVM RNA-class probability | 0.999819 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
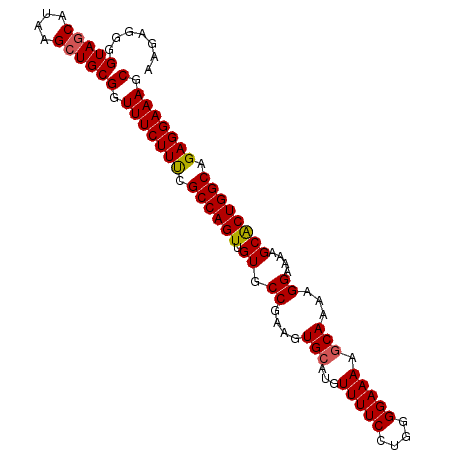
>3R_DroMel_CAF1 12495979 98 - 27905053 AAGAGGGGUAGCAUAAGCUGCGGUUUCUUUCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGC .......(((((....)))))(.((((((..((((((.((.((....(((...(((((....))))).)))...))....))))))))..)))))).) ( -34.50) >DroSec_CAF1 17473 98 - 1 CAGAGGGGUAGCAUAAGCUGCGGUUUCUUUCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGC .......(((((....)))))(.((((((..((((((.((.((....(((...(((((....))))).)))...))....))))))))..)))))).) ( -34.50) >DroSim_CAF1 16787 98 - 1 AAGAGGGGUACCAUAAGCUGCGGUUUCUUUCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGC ....(.(((.......))).)(.((((((..((((((.((.((....(((...(((((....))))).)))...))....))))))))..)))))).) ( -30.00) >DroEre_CAF1 17822 98 - 1 ACGAGGGGUAGCAUAAGCUGCGGUUUCUUCUGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAACCAAAAGGAAAAGCGCUGGCAGAGGAAAGC .......(((((....)))))(.((((((((((((((((..(....)..))..(((((((.((.....))...)))))))..)))))))))))))).) ( -42.70) >DroYak_CAF1 17751 98 - 1 AAGAGGGGUAGCAUAAGCUGCGGUUUCUUUUGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGC .......(((((....)))))(.((((((((((((((.((.((....(((...(((((....))))).)))...))....)))))))))))))))).) ( -38.10) >consensus AAGAGGGGUAGCAUAAGCUGCGGUUUCUUUCGCCAGUUGUGCCGAAGUGCAUGUUUUCCUGGGGAAAAGCAAAAGGAAAAGCACUGGCAGAGGAAAGC .......(((((....)))))(.(((((((.((((((.((.((....(((...(((((....))))).)))...))....)))))))).))))))).) (-32.54 = -32.62 + 0.08)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:38:19 2006