
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,337,858 – 12,337,999 |
| Length | 141 |
| Max. P | 0.995841 |

| Location | 12,337,858 – 12,337,961 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 89.26 |
| Mean single sequence MFE | -22.02 |
| Consensus MFE | -17.08 |
| Energy contribution | -17.48 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.31 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.62 |
| SVM RNA-class probability | 0.995841 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12337858 103 + 27905053 AUUAU-CCCACCUUUGCUAUAGCACUUAUGUAUCAUUAAGAAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAUGGGCUA .....-((((....(((....)))...........((((((((...........))))))))((...(((((((.........)))))))...))..))))... ( -16.80) >DroSec_CAF1 6749 104 + 1 AUUUUCCCCACCUCUGCUAUAUCACUUAUAUGUCAUUAAGAAAUUCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAGGGGUCA .........((((((((..(((.(((((...((.(((((((.....))))))).))...))))).)))((((((.........))))))....).))))))).. ( -19.80) >DroSim_CAF1 5881 103 + 1 AUUAU-CCCACCUCUGCUAUAUCACUUACAUGUCAUUAAGAAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCGGCCAGGGGCCA .....-....((((((((.(((.(((((...((.(((((((.....))))))).))...))))).)))((((((.........))))))...)).))))))... ( -19.30) >DroEre_CAF1 6208 103 + 1 AUUAU-GCCGCCCCUGCAAUAGCACUUAUGUAUCAUUAAGAAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAGGGGCUA .....-...((((((((((((((((....))....((((((((...........)))))))))))))(((((((.........)))))))..)).))))))).. ( -27.70) >DroYak_CAF1 2242 103 + 1 AUAAU-CCCACCCCGGCAAUAGCACUUAUGAAUCAUAAAGAAUUCCUUUUAAUUAUUCUUGAGCUAUAGUUUUCAACUUUUCUGAAAACUUCCGCCAGGGGCUA .....-....(((((((.(((((...((((...))))((((((...........))))))..)))))((((((((.......))))))))...))).))))... ( -26.50) >consensus AUUAU_CCCACCUCUGCUAUAGCACUUAUGUAUCAUUAAGAAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAGGGGCUA ..........((((((...................((((((((...........))))))))((...(((((((.........)))))))...))))))))... (-17.08 = -17.48 + 0.40)


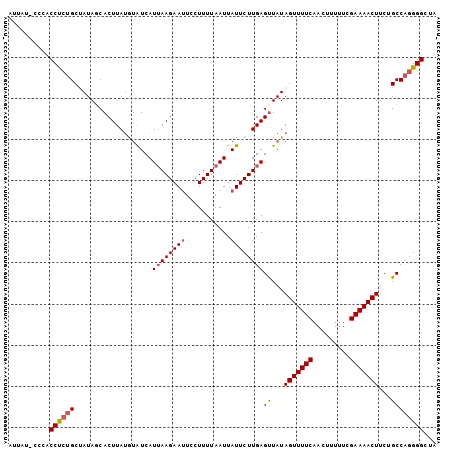
| Location | 12,337,858 – 12,337,961 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 89.26 |
| Mean single sequence MFE | -23.10 |
| Consensus MFE | -15.34 |
| Energy contribution | -15.18 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.506536 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12337858 103 - 27905053 UAGCCCAUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUUCUUAAUGAUACAUAAGUGCUAUAGCAAAGGUGGG-AUAAU ...(((((((((..(((((((((.....))))))))).......((((((...........)))))).............)))))).((....)))))-..... ( -19.00) >DroSec_CAF1 6749 104 - 1 UGACCCCUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGAAUUUCUUAAUGACAUAUAAGUGAUAUAGCAGAGGUGGGGAAAAU ...((((.......(((((((((.....)))))))))....((((((((.............)))))..((..((((.....))))..)))))..))))..... ( -20.62) >DroSim_CAF1 5881 103 - 1 UGGCCCCUGGCCGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUUCUUAAUGACAUGUAAGUGAUAUAGCAGAGGUGGG-AUAAU .((((...))))..(((((((((.....)))))))))....(((((((((...........))))))..((..((((.....))))..))))).....-..... ( -19.20) >DroEre_CAF1 6208 103 - 1 UAGCCCCUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUUCUUAAUGAUACAUAAGUGCUAUUGCAGGGGCGGC-AUAAU ..(((((((.(((.(((((((((.....))))))))).......((((((...........))))))..................))))))))))...-..... ( -26.80) >DroYak_CAF1 2242 103 - 1 UAGCCCCUGGCGGAAGUUUUCAGAAAAGUUGAAAACUAUAGCUCAAGAAUAAUUAAAAGGAAUUCUUUAUGAUUCAUAAGUGCUAUUGCCGGGGUGGG-AUUAU ..(((((.(((((.(((((((((.....))))))))).((((........(((((.((((....)))).))))).......))))))))))))))...-..... ( -29.86) >consensus UAGCCCCUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUUCUUAAUGAUACAUAAGUGCUAUAGCAGAGGUGGG_AUAAU ...(((((.((...(((((((((.....))))))))).......((((((...........))))))....................))..))).))....... (-15.34 = -15.18 + -0.16)



| Location | 12,337,897 – 12,337,999 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -21.10 |
| Consensus MFE | -19.18 |
| Energy contribution | -19.06 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.80 |
| SVM RNA-class probability | 0.977600 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12337897 102 + 27905053 AAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAUGGGCUACCAUAUUGGUCAACUUUGGCAUACUUUUAAGGCUGUUG ................(((((((....(((((((.........)))))))..(((((.((...(((.....)))...)).)))))....)))))))...... ( -19.20) >DroSec_CAF1 6789 102 + 1 AAAUUCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAGGGGUCACCAUAUUGGUCAACUUUGGCAUACUUUUAAAGCGGUUG .(((..((((((...............(((((((.........)))))))..((((((((...(((.....)))...)))))))).....))))))..))). ( -21.10) >DroSim_CAF1 5920 102 + 1 AAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCGGCCAGGGGCCACCAUAUUGGUCAACUUUGGCAUACUUUUAAAGCGGUUG ....((((((((...............(((((((.........)))))))...(((((((...(((.....)))...)))))))......)))))).))... ( -20.70) >DroEre_CAF1 6247 102 + 1 AAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAGGGGCUACCAUAUUGGUCAACUUUGGCAUACUUUUAAAGCUGUUG .....................((((..(((((((.........)))))))..((((((((...(((.....)))...)))))))).........)))).... ( -20.40) >DroYak_CAF1 2281 102 + 1 AAUUCCUUUUAAUUAUUCUUGAGCUAUAGUUUUCAACUUUUCUGAAAACUUCCGCCAGGGGCUACCAUAUUGGUCAACUUUGGCAUACUUUUAAAGCUGUUG .....................((((..((((((((.......))))))))...(((((((...(((.....)))...)))))))..........)))).... ( -24.10) >consensus AAUUCCUUUUAAUUAUUCUUGAGUUAUAGUUUUCAACUUUUUCGAAAACUUCUGCCAGGGGCUACCAUAUUGGUCAACUUUGGCAUACUUUUAAAGCUGUUG ......................(((..(((((((.........)))))))...(((((((...(((.....)))...)))))))..........)))..... (-19.18 = -19.06 + -0.12)



| Location | 12,337,897 – 12,337,999 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -20.38 |
| Consensus MFE | -18.42 |
| Energy contribution | -17.82 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.44 |
| SVM RNA-class probability | 0.993960 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12337897 102 - 27905053 CAACAGCCUUAAAAGUAUGCCAAAGUUGACCAAUAUGGUAGCCCAUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUU ......((((.......(((((..((((.((.....))))))...)))))..(((((((((.....)))))))))...................)))).... ( -21.60) >DroSec_CAF1 6789 102 - 1 CAACCGCUUUAAAAGUAUGCCAAAGUUGACCAAUAUGGUGACCCCUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGAAUUU ......((((.......(((((..(((.(((.....))))))...)))))..(((((((((.....)))))))))..................))))..... ( -19.30) >DroSim_CAF1 5920 102 - 1 CAACCGCUUUAAAAGUAUGCCAAAGUUGACCAAUAUGGUGGCCCCUGGCCGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUU ...((..(((((...(((.....((((((((.....)))((((...))))..(((((((((.....))))))))).))))).....))).))))).)).... ( -21.50) >DroEre_CAF1 6247 102 - 1 CAACAGCUUUAAAAGUAUGCCAAAGUUGACCAAUAUGGUAGCCCCUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUU ......((((.......(((((..((((.((.....))))))...)))))..(((((((((.....)))))))))..................))))..... ( -19.40) >DroYak_CAF1 2281 102 - 1 CAACAGCUUUAAAAGUAUGCCAAAGUUGACCAAUAUGGUAGCCCCUGGCGGAAGUUUUCAGAAAAGUUGAAAACUAUAGCUCAAGAAUAAUUAAAAGGAAUU ....((((.........(((((..((((.((.....))))))...)))))..(((((((((.....)))))))))..))))..................... ( -20.10) >consensus CAACAGCUUUAAAAGUAUGCCAAAGUUGACCAAUAUGGUAGCCCCUGGCAGAAGUUUUCGAAAAAGUUGAAAACUAUAACUCAAGAAUAAUUAAAAGGAAUU ......((((.......(((((..(((.(((.....))))))...)))))..(((((((((.....)))))))))...................)))).... (-18.42 = -17.82 + -0.60)


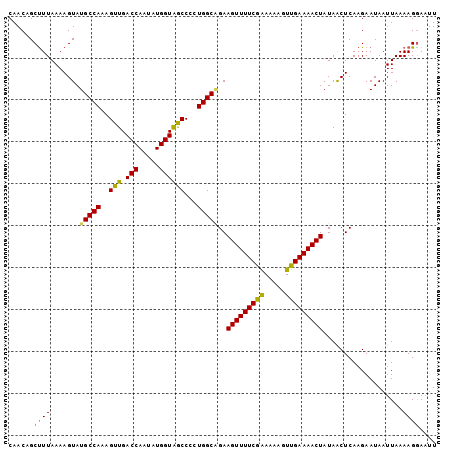
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:36:30 2006