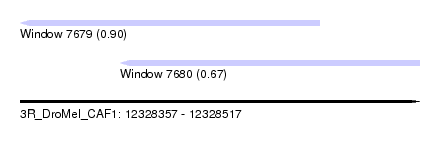
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,328,357 – 12,328,517 |
| Length | 160 |
| Max. P | 0.898492 |

| Location | 12,328,357 – 12,328,477 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.56 |
| Mean single sequence MFE | -39.53 |
| Consensus MFE | -31.63 |
| Energy contribution | -30.53 |
| Covariance contribution | -1.10 |
| Combinations/Pair | 1.16 |
| Mean z-score | -3.09 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.898492 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12328357 120 - 27905053 UUUGAAUUCGGUUUCAGCUCGAGCGGGUUCAUGUUCGAAGCGAGCAAGUUCGAGUUCGUAUCGAACUUAAGAGAGCCAACUAAAAGAGUGGGCUAAGAGUUUAAGGAACUGUUUAUAUAU ........(((((((.(((((((((......)))))).)))((((((((((((.......)))))))).....((((.(((.....))).))))....))))..)))))))......... ( -37.00) >DroSec_CAF1 19502 120 - 1 UUUGAAUUUGGUUUCGGCUCGAGCGGGCUCAUGUUCGAAGCGAGCCGGUUCGAGUUCGAAUCGAACUUAAGAGAGCAAACUAAAAGAGUGGGUGAAGAGUUUGAAGAACAGUUUCUAUAC (((((((((((..((((((((..(((((....)))))...)))))))).)))))))))))((((((((......((..(((.....)))..))...))))))))((((....)))).... ( -41.50) >DroSim_CAF1 15534 120 - 1 UUUGAAUUUGGUGCCAGCUCGAGCGAGCUCAUGUUCGAAGCGAGCCGGUUCGAGUUCGAAUCGAACUUAAGAGAGCAAACUAAAAGAGUGGGUGAAGAGUUUGAAGAACAGUUUCUAUAC (((((((((((.(((.(((((..(((((....)))))...))))).))))))))))))))((((((((......((..(((.....)))..))...))))))))((((....)))).... ( -40.10) >consensus UUUGAAUUUGGUUUCAGCUCGAGCGGGCUCAUGUUCGAAGCGAGCCGGUUCGAGUUCGAAUCGAACUUAAGAGAGCAAACUAAAAGAGUGGGUGAAGAGUUUGAAGAACAGUUUCUAUAC (((((((((....((.((((.((...((((..((((((..(((((........)))))..))))))......))))...))....)))).))....)))))))))............... (-31.63 = -30.53 + -1.10)


| Location | 12,328,397 – 12,328,517 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -41.43 |
| Consensus MFE | -36.48 |
| Energy contribution | -35.93 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.665474 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12328397 120 - 27905053 UGGGCAACAUAUACAACGCUUUUGGCGUCGCUGUUUUGGUUUUGAAUUCGGUUUCAGCUCGAGCGGGUUCAUGUUCGAAGCGAGCAAGUUCGAGUUCGUAUCGAACUUAAGAGAGCCAAC ..(((..........((((.....)))).((((((((((...((((((((.((.......)).))))))))...))))))).)))((((((((.......))))))))......)))... ( -38.10) >DroSec_CAF1 19542 120 - 1 UGGGCAACAUAUACAACGCUUUUGGCGUCGCUGUUUUGGUUUUGAAUUUGGUUUCGGCUCGAGCGGGCUCAUGUUCGAAGCGAGCCGGUUCGAGUUCGAAUCGAACUUAAGAGAGCAAAC ..(....).......((((.....)))).(((.((((((((((((((((((..((((((((..(((((....)))))...)))))))).)))))))))....)))).))))).))).... ( -43.80) >DroSim_CAF1 15574 120 - 1 UGGGCAACAUAUACAACGCUUUUGGCGUCGCUGUUUUGGUUUUGAAUUUGGUGCCAGCUCGAGCGAGCUCAUGUUCGAAGCGAGCCGGUUCGAGUUCGAAUCGAACUUAAGAGAGCAAAC ..(....).......((((.....)))).(((.((((((((((((((((((.(((.(((((..(((((....)))))...))))).))))))))))))....)))).))))).))).... ( -42.40) >consensus UGGGCAACAUAUACAACGCUUUUGGCGUCGCUGUUUUGGUUUUGAAUUUGGUUUCAGCUCGAGCGGGCUCAUGUUCGAAGCGAGCCGGUUCGAGUUCGAAUCGAACUUAAGAGAGCAAAC ..(....).......((((.....)))).(((.((((((((((((((((((..((.(((((..(((((....)))))...))))).)).)))))))))....)))).))))).))).... (-36.48 = -35.93 + -0.55)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:36:12 2006