
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,180,345 – 12,180,467 |
| Length | 122 |
| Max. P | 0.961480 |

| Location | 12,180,345 – 12,180,438 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 80.99 |
| Mean single sequence MFE | -28.50 |
| Consensus MFE | -20.38 |
| Energy contribution | -21.75 |
| Covariance contribution | 1.37 |
| Combinations/Pair | 1.40 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.937485 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12180345 93 + 27905053 AUACUUUUUUGUGUGCGGCUAUUAUCUGUUGUCGUGG---GCCGCAAAACGGAGCCAAAGCCAAUUAUUCUCGGCUUUGGCUGUUGUCGACUUUAU ((((......))))((((((.....(....).....)---)))))...((((((((((((((..........)))))))))).))))......... ( -30.70) >DroPse_CAF1 6586 94 + 1 AUACAUGAUUGUUUUUGGCAUUUAUCUGGUAUCUGGUACUGCCGCC-AGCGAAGCCGACGCCAAUUAUUUUCAACUCUU-UUGUUGUUCACUUUGU ....(((((((...(((((.(((..(((((....((.....)))))-)).))))))))...)))))))...((((....-..)))).......... ( -19.90) >DroSec_CAF1 5024 93 + 1 AUACUUUUUUGUUUGCGGCUAUCAUCUGUUGUCGUGG---GCCGCAAAACGAAGCCAAAGCCAAUUAUUCUCGGCUUUGGCUGUUGUCGACUUUAU ...........(((((((((..((.....)).....)---))))))))((((((((((((((..........)))))))))).))))......... ( -32.40) >DroSim_CAF1 5095 93 + 1 AUACUUUUUUGUUUGCGGCUAUCAUCUGUUGUCGUGG---GCCGCAAAACGAAGCCAAAGCCAAUUAUUCUCGGCUUUGGCUGUUGUUGACUUUAU ............((((((((..((.....)).....)---)))))))(((((((((((((((..........)))))))))).)))))........ ( -33.00) >DroEre_CAF1 4313 93 + 1 GUACUAUAUUGUUUGCGGCUAUCAUCUGCUGUCAUGG---GCCACAAAACAGAGCCAAAGCCAAUUAUACUCGGCUUUGGCUUUAGUCGACUUAAU (.((((...((((((.((((.....(....).....)---))).).)))))(((((((((((..........))))))))))))))))........ ( -28.40) >DroYak_CAF1 4929 93 + 1 AAACUAUACGGUUUGCGGCUAUCAACUGCUGUCGUGG---GCCACAAAGCGGACCCAAAGUCAAUUAUUCUCGGCUUUGGCUGUUGUCGACUUUAU .........((((..((((...........))))..)---)))....(((((..((((((((..........)))))))))))))........... ( -26.60) >consensus AUACUUUAUUGUUUGCGGCUAUCAUCUGUUGUCGUGG___GCCGCAAAACGAAGCCAAAGCCAAUUAUUCUCGGCUUUGGCUGUUGUCGACUUUAU ........(((((((((((..((((........))))...))))).))))))((((((((((..........)))))))))).............. (-20.38 = -21.75 + 1.37)


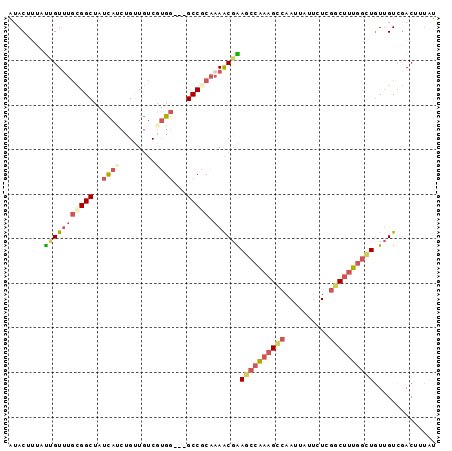
| Location | 12,180,345 – 12,180,438 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 80.99 |
| Mean single sequence MFE | -28.34 |
| Consensus MFE | -19.87 |
| Energy contribution | -21.32 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.19 |
| Mean z-score | -3.50 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961480 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12180345 93 - 27905053 AUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUCCGUUUUGCGGC---CCACGACAACAGAUAAUAGCCGCACACAAAAAAGUAU ........(((...((((((((((((......))))))))))))..))).((((((---..................))))))............. ( -28.97) >DroPse_CAF1 6586 94 - 1 ACAAAGUGAACAACAA-AAGAGUUGAAAAUAAUUGGCGUCGGCUUCGCU-GGCGGCAGUACCAGAUACCAGAUAAAUGCCAAAAACAAUCAUGUAU .....((((.((((..-....)))).......(((((((((((...)))-)))((.....))...............)))))......)))).... ( -20.70) >DroSec_CAF1 5024 93 - 1 AUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUUCGUUUUGCGGC---CCACGACAACAGAUGAUAGCCGCAAACAAAAAAGUAU .......(((....((((((((((((......))))))))))))))).((((((((---...((.(....).))...))))))))........... ( -31.60) >DroSim_CAF1 5095 93 - 1 AUAAAGUCAACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUUCGUUUUGCGGC---CCACGACAACAGAUGAUAGCCGCAAACAAAAAAGUAU ..............((((((((((((......))))))))))))..((((.(((((---...((.(....).))...))))))))).......... ( -30.80) >DroEre_CAF1 4313 93 - 1 AUUAAGUCGACUAAAGCCAAAGCCGAGUAUAAUUGGCUUUGGCUCUGUUUUGUGGC---CCAUGACAGCAGAUGAUAGCCGCAAACAAUAUAGUAC .........((((.((((((((((((......)))))))))))).(((((.(((((---.(((........)))...))))))))))...)))).. ( -33.00) >DroYak_CAF1 4929 93 - 1 AUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGACUUUGGGUCCGCUUUGUGGC---CCACGACAGCAGUUGAUAGCCGCAAACCGUAUAGUUU ........(((.....((((((.(((......))).))))))...((.((((((((---...((((....))))...)))))))).))....))). ( -25.00) >consensus AUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUCCGUUUUGCGGC___CCACGACAACAGAUGAUAGCCGCAAACAAAAAAGUAU ..............((((((((((((......))))))))))))..(.((((((((.....................))))))))).......... (-19.87 = -21.32 + 1.45)



| Location | 12,180,365 – 12,180,467 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 82.43 |
| Mean single sequence MFE | -35.88 |
| Consensus MFE | -19.24 |
| Energy contribution | -19.85 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.32 |
| Mean z-score | -3.19 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.813454 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12180365 102 - 27905053 -GCAGCA-GGGUCACAAAAGUUCGAA--UGGCGAUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUCCGUUUUGCGGC---CCACGACAACAGAUAAU -......-(((((.(((((.......--((((......)))).....((((((((((((......))))))))))))...))))).)))---))............... ( -36.20) >DroPse_CAF1 6606 107 - 1 CGUAGAAGGGGUCGCAAAAGUUCGCCCCCCGCGACAAAGUGAACAACAA-AAGAGUUGAAAAUAAUUGGCGUCGGCUUCGCU-GGCGGCAGUACCAGAUACCAGAUAAA .......(((((.((....))..))))).(((......)))..((((..-....))))......((((.(((((((...)))-)))).))))................. ( -27.70) >DroSec_CAF1 5044 102 - 1 -GCAGCG-GGGUCACAAAAGUUCGAA--UGGCGAUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUUCGUUUUGCGGC---CCACGACAACAGAUGAU -...(((-(((((.(((((...((((--((((......))))......(((((((((((......)))))))))))))))))))).)))---)).)).).......... ( -38.80) >DroSim_CAF1 5115 102 - 1 -GCAGCA-GGGUCACAAAAGUUCGAA--UGGCGAUAAAGUCAACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUUCGUUUUGCGGC---CCACGACAACAGAUGAU -......-(((((.(((((...((((--((((......))))......(((((((((((......)))))))))))))))))))).)))---))............... ( -38.00) >DroEre_CAF1 4333 102 - 1 -GCAGCA-GGGUCACAAAAGUUCGAG--UAGCGAUUAAGUCGACUAAAGCCAAAGCCGAGUAUAAUUGGCUUUGGCUCUGUUUUGUGGC---CCAUGACAGCAGAUGAU -((..((-(((((((((((.......--(((((((...)))).))).((((((((((((......))))))))))))...)))))))))---)).))...))....... ( -43.20) >DroYak_CAF1 4949 102 - 1 -GCAGCA-GGGUCACAAAAGUUCGAA--CUGCGAUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGACUUUGGGUCCGCUUUGUGGC---CCACGACAGCAGUUGAU -......-((((((((((.((((((.--((.......))))))......((((((.(((......))).))))))....))))))))))---)).((((....)))).. ( -31.40) >consensus _GCAGCA_GGGUCACAAAAGUUCGAA__UGGCGAUAAAGUCGACAACAGCCAAAGCCGAGAAUAAUUGGCUUUGGCUCCGUUUUGCGGC___CCACGACAACAGAUGAU ..........(((........(((.......)))....((((.....((((((((((((......))))))))))))(((.....))).......))))....)))... (-19.24 = -19.85 + 0.61)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:35:08 2006