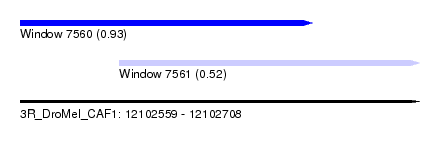

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,102,559 – 12,102,708 |
| Length | 149 |
| Max. P | 0.926105 |

| Location | 12,102,559 – 12,102,668 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 95.75 |
| Mean single sequence MFE | -16.38 |
| Consensus MFE | -14.66 |
| Energy contribution | -14.66 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.926105 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12102559 109 + 27905053 GCAACUAACCCAGCUAAACUAACGUAAUUUUGUUGCUAGCCAACAAAACAAACUUACUCACACCCAUCUAG----AUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGA .........((((((((((....))..((((((((.....)))))))).......................----.....................))))))))......... ( -16.00) >DroSec_CAF1 8374 109 + 1 GCAACUAACCCUGCUAGACUAACGUAAUUUUGUUGCUAGCCAACAAAACAAACUUACUCACACCCAUCUAG----AUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGA ((...........((((((....))..((((((((.....))))))))...................))))----.........((((((.((......))))))))..)).. ( -17.20) >DroSim_CAF1 8370 109 + 1 GCAACUAACCCAGCUAAACUAACGUAAUUUUGUUGCUAGCCAACAAAACAAACUUACUCACACCCAUCUAG----AUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGA .........((((((((((....))..((((((((.....)))))))).......................----.....................))))))))......... ( -16.00) >DroEre_CAF1 8267 113 + 1 GCAACUAACCCAGCUAAACUAACGUAAUUUUGUUGCUAGCCAACAAAACAAACUUACUCACACCCAUCUAUACAUAUAUUCAUAUUUCCACACAUUUUAGUUGGAGACAGCGA .........((((((((((....))..((((((((.....))))))))................................................))))))))......... ( -16.00) >DroYak_CAF1 8313 109 + 1 GGAACUAAGCCAGCUAAACUAACGUAAUUUUGUUGCUAGCCAACAAAACAAACUUACUCACACCCAUCUCC----AUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGA ((((.............((....))..((((((((.....)))))))).......................----..........))))..........((((....)))).. ( -16.70) >consensus GCAACUAACCCAGCUAAACUAACGUAAUUUUGUUGCUAGCCAACAAAACAAACUUACUCACACCCAUCUAG____AUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGA ............(((........((((((((((((.....)))))))).....))))...........................((((((.((......)))))))).))).. (-14.66 = -14.66 + 0.00)



| Location | 12,102,596 – 12,102,708 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 97.62 |
| Mean single sequence MFE | -12.67 |
| Consensus MFE | -12.22 |
| Energy contribution | -12.00 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.96 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.517713 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12102596 112 + 27905053 AGCCAACAAAACAAACUUACUCACACCCAUCUAGAUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGAAUGUAAUAAAAAACAAACUUUUUGUAUUACUUUUUAUUAU ..................................((((((.(.((((((.((......))))))))..).))))))((((((((((((.....)))).....)))))))).. ( -12.20) >DroSim_CAF1 8407 112 + 1 AGCCAACAAAACAAACUUACUCACACCCAUCUAGAUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGAAUGUAAUAAAAAACAAACUUUUUGUAUUACUUUUUAUUAU ..................................((((((.(.((((((.((......))))))))..).))))))((((((((((((.....)))).....)))))))).. ( -12.20) >DroYak_CAF1 8350 112 + 1 AGCCAACAAAACAAACUUACUCACACCCAUCUCCAUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGAAUGUAACAAAAAACAAACUUUUUGUUUUACUUUUUAUUAU ..................................((((((.(.((((((.((......))))))))..).))))))((((((((......)))))))).............. ( -13.60) >consensus AGCCAACAAAACAAACUUACUCACACCCAUCUAGAUAUUCACAUUUCCACACAUUUUAGUUGGAGACAGCGAAUGUAAUAAAAAACAAACUUUUUGUAUUACUUUUUAUUAU ..................................((((((.(.((((((.((......))))))))..).)))))).(((((((......)))))))............... (-12.22 = -12.00 + -0.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:34:17 2006