
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,082,775 – 12,082,975 |
| Length | 200 |
| Max. P | 0.997667 |

| Location | 12,082,775 – 12,082,895 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.42 |
| Mean single sequence MFE | -43.28 |
| Consensus MFE | -36.46 |
| Energy contribution | -36.82 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.788971 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12082775 120 - 27905053 CACCUCUAACCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUUGGCGGGAGUGGCACCUCCGCUUGACUUUUCAUUGCUGCGACGGCGGUAGUAACUUCAUGACCAUUGCUAAU ..((.(((((..((((((((((.....))))))))))((....)).))))).))((((((.....))))))........((((((((....))))))))..................... ( -40.00) >DroSec_CAF1 19472 120 - 1 CACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUCGGCGGCAGUGGCACCUCCGCUUGACUUUUCAUUGCUGCGACGGCGGUAGUAACUUCAUGACCAUUGCUAAU ......((((..((((((((((.....))))))))))((((.((((((((.(((((((((....((.....))....))))))))))))))))))).))...............)))).. ( -47.50) >DroSim_CAF1 19660 120 - 1 CACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGACUUUUCAUUGCUGCGACGGCGGUAGUAACUUCAUGACCAUUGCUAAU ......((((..((((((((((.....))))))))))((((.((((((((.(((((((((.....))))))((....))....))))))))))))).))...............)))).. ( -44.20) >DroEre_CAF1 21566 120 - 1 CACCUCUAGCCUAGCGAUACUGUCCUCCGGGAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGGCUUUUUAGGGCUGCGACGGCGGUAGUAACUUCAUGACCAUUGCUAAU ..(((..((((.((((.....((((....)))).(((((.(((((.....))))).))))).....)))).))))....)))(((((....)))))(((((...........)))))... ( -43.20) >DroYak_CAF1 21951 120 - 1 CACCUCUAGUCUAGCGAUACUGUCCUCUGGUAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGGCUUUUAAGGGCUGCGACGGCGGUAGUAACUUCAUGACCAUUGCUAAU ..(((..((((.(((((((((.......))))))(((((.(((((.....))))).)))))......))).))))....)))(((((....)))))(((((...........)))))... ( -41.50) >consensus CACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGACUUUUCAUUGCUGCGACGGCGGUAGUAACUUCAUGACCAUUGCUAAU ...........(((((((...(((...(((....(((((.(((((.....))))).)))))....)))...))).....((((((((....))))))))............))))))).. (-36.46 = -36.82 + 0.36)

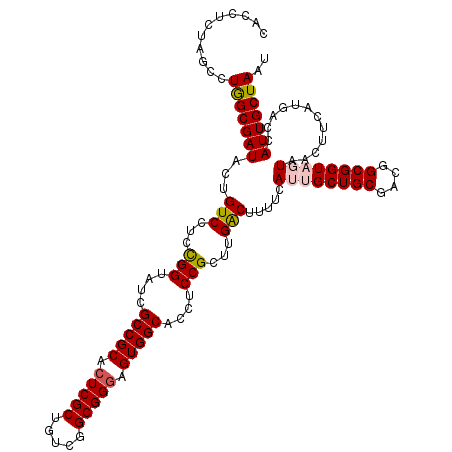

| Location | 12,082,815 – 12,082,935 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.25 |
| Mean single sequence MFE | -48.90 |
| Consensus MFE | -45.60 |
| Energy contribution | -46.32 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.91 |
| SVM RNA-class probability | 0.997667 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12082815 120 - 27905053 CAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAACCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUUGGCGGGAGUGGCACCUCCGCUUGACUUUUCA ....((((.....))))....((((...(((.((((((.((((.........((((((((((.....))))))))))...(((((.....))))).)))))))))).)))...))))... ( -50.30) >DroSec_CAF1 19512 120 - 1 CAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUCGGCGGCAGUGGCACCUCCGCUUGACUUUUCA ((((((.((((.(.(((...........(((((((((....))))).)))).((((((((((.....))))))))))......))).).))))))))))((......))........... ( -50.10) >DroSim_CAF1 19700 120 - 1 CAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGACUUUUCA ....((((.....))))....((((...(((.((((((.((((.(((.(((.((((((((((.....))))))))))((....))....))).))))))))))))).)))...))))... ( -52.80) >DroEre_CAF1 21606 120 - 1 CAUUGUUCGUCCAUGGCUGACAAGGAACGGCUGAGGUGACCACCUCUAGCCUAGCGAUACUGUCCUCCGGGAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGGCUUUUUA ......((((((.((.....)).)).))))..(((((....))))).((((.((((.....((((....)))).(((((.(((((.....))))).))))).....)))).))))..... ( -45.90) >DroYak_CAF1 21991 120 - 1 CAUUGCUCGUCCAGGGCUGACAAGGAACGGCUGAGGUGACCACCUCUAGUCUAGCGAUACUGUCCUCUGGUAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGGCUUUUAA ....((((.....))))....((((...(((.((((((.((((..........((((((((.......))))))))....(((((.....))))).)))))))))).)))...))))... ( -45.40) >consensus CAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCACUCGCUGUCGGCGGGAGUGGCACCUCCGCUUGACUUUUCA ....((((.....))))....((((...(((.((((((.((((.(((.(((.((((((((((.....))))))))))((....))....))).))))))))))))).)))...))))... (-45.60 = -46.32 + 0.72)



| Location | 12,082,855 – 12,082,975 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.00 |
| Mean single sequence MFE | -54.08 |
| Consensus MFE | -46.12 |
| Energy contribution | -47.28 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989305 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12082855 120 - 27905053 GAUGCUGCGGGUCCUGCCACCGCGGAAGGAGUAGUGGGCCCAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAACCUGGCGAUACUGUCCUCCGGUAUCGCCGCA ..(((((.(((((((((....))(((..((((((((....)))))))).)))))))))).))......((..(((((....)))))...)).((((((((((.....))))))))))))) ( -55.00) >DroSec_CAF1 19552 120 - 1 GAUGCAGCGGGUCCUGUCACCGCGGAAGGAGCAGUGGGCCCAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCA ..(((.(.((((((((......(....)((((((((....))))))))...)))))))).).......(((((((((....))))).)))).((((((((((.....))))))))))))) ( -61.10) >DroSim_CAF1 19740 120 - 1 GAUGCAGCGGGUCCUGCCACCGCGGAAGGAGUAGUGGGCCCAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCA ..(((.(.(((((((((....))(((..((((((((....)))))))).)))))))))).).......(((((((((....))))).)))).((((((((((.....))))))))))))) ( -60.30) >DroEre_CAF1 21646 120 - 1 GAUCCAGCAGGUCCUGCCACCGCCGAAGGAGUCGUGGGCGCAUUGUUCGUCCAUGGCUGACAAGGAACGGCUGAGGUGACCACCUCUAGCCUAGCGAUACUGUCCUCCGGGAUCGCCGCA (((((.(.(((((((....((......))(((((((((((.......)))))))))))....))))..(((((((((....))))).))))............))).).)))))...... ( -46.90) >DroYak_CAF1 22031 120 - 1 GAUCCAGCGGGUCCUGCCACCGCCGAUGGAGUCGUGGACGCAUUGCUCGUCCAGGGCUGACAAGGAACGGCUGAGGUGACCACCUCUAGUCUAGCGAUACUGUCCUCUGGUAUCGCCGCA ..(((.((((.........))))...((.((((.((((((.......)))))).))))..)).)))..(((((((((....))))).))))..((((((((.......)))))))).... ( -47.10) >consensus GAUGCAGCGGGUCCUGCCACCGCGGAAGGAGUAGUGGGCCCAUUGCUCGUCCAGGGCUCACAAGGAACGGCUGAGGUGACCACCUCUAGCCUGGCGAUACUGUCCUCCGGUAUCGCCGCA ......((((((((((......(....)((((((((....))))))))...)))))))).........(((((((((....))))).)))).((((((((((.....)))))))))))). (-46.12 = -47.28 + 1.16)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:33:54 2006