
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,080,021 – 12,080,135 |
| Length | 114 |
| Max. P | 0.999397 |

| Location | 12,080,021 – 12,080,135 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.45 |
| Mean single sequence MFE | -39.44 |
| Consensus MFE | -36.16 |
| Energy contribution | -36.84 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.92 |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.989873 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12080021 114 + 27905053 ACUGGCUUUUGUGUUGCUGCUGCGGUGCUUGGUUGCU------GUUGCCGCUUGAAACUGCCUGAGUGAUGUUGGUGUGGCAAUAAAAUUCAACUUCCAGCAACAUAGCCAGCGAACGCA .((((((...(((((((((..(((((....(((.((.------...)).)))....))))).(((((..(((((......)))))..))))).....)))))))))))))))........ ( -40.00) >DroSec_CAF1 16764 114 + 1 ACUGGCUUUUGUGUUGCUGCUGCUGUGCUUGGUUGCU------GUUGCCGCUUGAAACUGCCUGAGUGAUGUUGGUGUGGCAAUAAAAUUCAACUUCCAGCAACAUAGCCAGCGAACGCA .((((((...(((((((((.......((......))(------(((((((((..(.(((.....)))....)..).)))))))))............)))))))))))))))........ ( -37.80) >DroSim_CAF1 16956 114 + 1 ACUGGCUUUUGUGUUGCUGCUGCGGUGCUUGGUUGCU------GUUGCCGCUUGAAACUGCCUGAGUGAUGUUGGUGUGGCAAUAAAAUUCAACUUCCAGCAACAUAGCCAGCGAACGCA .((((((...(((((((((..(((((....(((.((.------...)).)))....))))).(((((..(((((......)))))..))))).....)))))))))))))))........ ( -40.00) >DroEre_CAF1 18799 120 + 1 ACUGGCUUUUGUAUUGCUGCUGCAGUGCUUGGUUGCUGUUGCUGUUGCCGCUUGAAACUGCCUGAGUGAUGUUGGUGUGGCAAUAAAAUUCAACUUCCAGCAACAUUGCCAGCGAACGCA .(((((....(((((((....)))))))...((((((((((.((((((((((..(.(((.....)))....)..).))))))))).....))))....))))))...)))))........ ( -39.90) >DroYak_CAF1 19156 114 + 1 ACUGGCUUUUGUGUUGCUGCUGCGGUGCCUGGUCGCU------GUUGCCGCUUAAAACUGCCUGAGUGAUGUUGGUGUGGCAAUAAAAUUCAACUUCCAGCAACAUUGCCAGCGAACGCA .(((((....(((((((((..(((((....(((.((.------...)).)))....))))).(((((..(((((......)))))..))))).....))))))))).)))))........ ( -39.50) >consensus ACUGGCUUUUGUGUUGCUGCUGCGGUGCUUGGUUGCU______GUUGCCGCUUGAAACUGCCUGAGUGAUGUUGGUGUGGCAAUAAAAUUCAACUUCCAGCAACAUAGCCAGCGAACGCA .((((((...(((((((((..(((((....(((.((..........)).)))....))))).(((((..(((((......)))))..))))).....)))))))))))))))........ (-36.16 = -36.84 + 0.68)


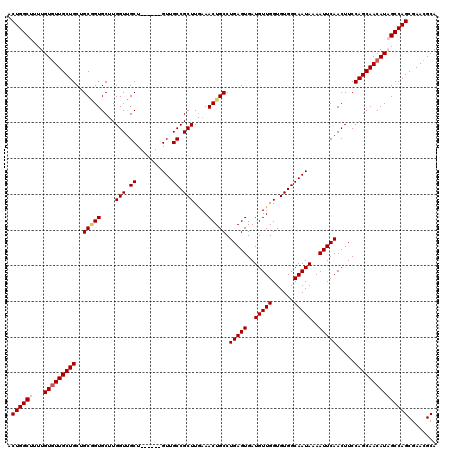
| Location | 12,080,021 – 12,080,135 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.45 |
| Mean single sequence MFE | -35.45 |
| Consensus MFE | -32.27 |
| Energy contribution | -32.51 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.57 |
| SVM RNA-class probability | 0.999397 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12080021 114 - 27905053 UGCGUUCGCUGGCUAUGUUGCUGGAAGUUGAAUUUUAUUGCCACACCAACAUCACUCAGGCAGUUUCAAGCGGCAAC------AGCAACCAAGCACCGCAGCAGCAACACAAAAGCCAGU .......(((((((.((((((((....(((((....((((((................)))))))))))((((....------.((......)).))))..))))))))....))))))) ( -36.39) >DroSec_CAF1 16764 114 - 1 UGCGUUCGCUGGCUAUGUUGCUGGAAGUUGAAUUUUAUUGCCACACCAACAUCACUCAGGCAGUUUCAAGCGGCAAC------AGCAACCAAGCACAGCAGCAGCAACACAAAAGCCAGU ((((((.(((((((..(((((((...((((......((((((................))))))......))))..)------))))))..))).))))))).))).............. ( -35.39) >DroSim_CAF1 16956 114 - 1 UGCGUUCGCUGGCUAUGUUGCUGGAAGUUGAAUUUUAUUGCCACACCAACAUCACUCAGGCAGUUUCAAGCGGCAAC------AGCAACCAAGCACCGCAGCAGCAACACAAAAGCCAGU .......(((((((.((((((((....(((((....((((((................)))))))))))((((....------.((......)).))))..))))))))....))))))) ( -36.39) >DroEre_CAF1 18799 120 - 1 UGCGUUCGCUGGCAAUGUUGCUGGAAGUUGAAUUUUAUUGCCACACCAACAUCACUCAGGCAGUUUCAAGCGGCAACAGCAACAGCAACCAAGCACUGCAGCAGCAAUACAAAAGCCAGU .......((((((..((((((((...((((......((((((................))))))......))))..))))))))((......))..(((....)))........)))))) ( -35.69) >DroYak_CAF1 19156 114 - 1 UGCGUUCGCUGGCAAUGUUGCUGGAAGUUGAAUUUUAUUGCCACACCAACAUCACUCAGGCAGUUUUAAGCGGCAAC------AGCGACCAGGCACCGCAGCAGCAACACAAAAGCCAGU .......((((((..((((((((...((((......((((((................))))))......))))...------.(((.........)))..)))))))).....)))))) ( -33.39) >consensus UGCGUUCGCUGGCUAUGUUGCUGGAAGUUGAAUUUUAUUGCCACACCAACAUCACUCAGGCAGUUUCAAGCGGCAAC______AGCAACCAAGCACCGCAGCAGCAACACAAAAGCCAGU .......(((((((.((((((((...((((......((((((................))))))......))))..........((......)).......))))))))....))))))) (-32.27 = -32.51 + 0.24)

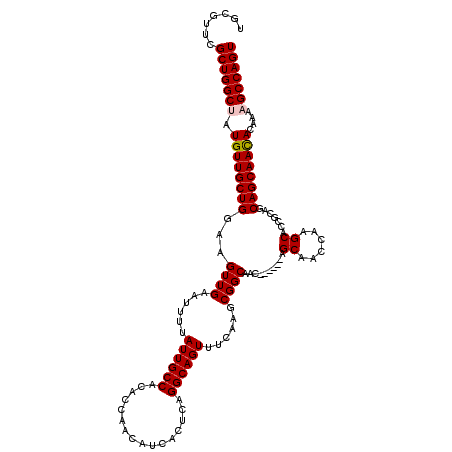

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:33:42 2006