
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 12,016,154 – 12,016,252 |
| Length | 98 |
| Max. P | 0.991868 |

| Location | 12,016,154 – 12,016,252 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 93.96 |
| Mean single sequence MFE | -29.54 |
| Consensus MFE | -21.94 |
| Energy contribution | -21.98 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.689376 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12016154 98 + 27905053 UCGAUGGGAGAAAGAUAUGGGAGUGCG--UAUAUGUGGCCAGAAAAUUAUGUAUACGCAGCGUUUCCUCAUUGAGAGGCGCUGCUGACGCAAAAUAAAUG .......................((((--(((((((((........))))))))))(((((((((((.....).))))))))))....)))......... ( -26.60) >DroSec_CAF1 121011 100 + 1 UCGAUGGGAGAAAGAUAUGGGAGUGCGUGUAUAUGUGGCCAGAAAAUUGUGUAUACGCAGCGUUUCCUCAUUGAGAGGCGCUGCUGACGCAAAAUAAAUG .......................((((((((((((..(........)..)))))))(((((((((((.....).))))))))))..)))))......... ( -29.10) >DroSim_CAF1 121293 100 + 1 UCGAUGGGAGAAAGAUAUGGGAGUGCGUGUAUAUGUGGCCAGAAAAUUAUGUAUACGCAGCGUUUCCUCAUUGAGAGGCGCUGCUGACGCAAAAUAAAUG .......................(((((((((((((((........))))))))))(((((((((((.....).))))))))))..)))))......... ( -29.50) >DroEre_CAF1 120720 98 + 1 UCCACGUCCGAAAGAUAUGAGAGUGCG--UAUAUGUGGCCAGAAAAUUAUGUAUACGCAGCGUUUCCUCAUUGUGAGGAGCUGCUGACGCAAAAUAAAUG ....(((((....)..........(((--(((((((((........))))))))))))((((.((((((.....)))))).))))))))........... ( -29.30) >DroYak_CAF1 123834 98 + 1 UCGACGUUCGAAAGAUAUGCGAGUGCG--UAUAUGUGGCCAGAAAAUUAUGUAUACGCAGCGUUUCCUCAUUGAGAGGCGCUGCUGACGCAAAAUAAAUG (((.(((((....)).)))))).((((--(((((((((........))))))))))(((((((((((.....).))))))))))....)))......... ( -33.20) >consensus UCGAUGGGAGAAAGAUAUGGGAGUGCG__UAUAUGUGGCCAGAAAAUUAUGUAUACGCAGCGUUUCCUCAUUGAGAGGCGCUGCUGACGCAAAAUAAAUG .......................((((..((((..(((........)))..)))).(((((((((((.....).))))))))))...))))......... (-21.94 = -21.98 + 0.04)

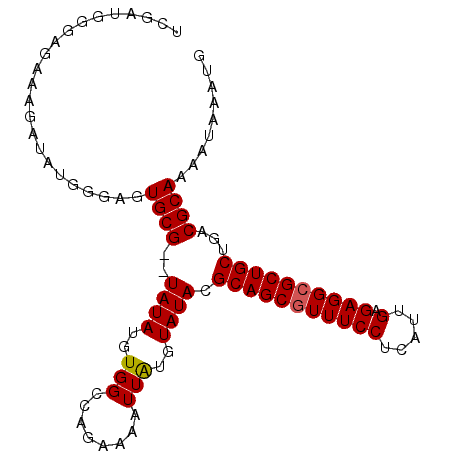
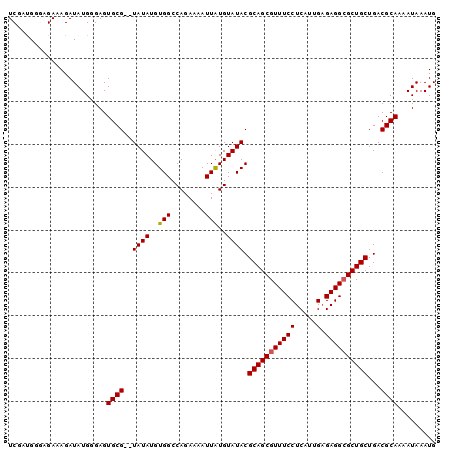
| Location | 12,016,154 – 12,016,252 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 93.96 |
| Mean single sequence MFE | -23.02 |
| Consensus MFE | -19.16 |
| Energy contribution | -19.36 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.29 |
| SVM RNA-class probability | 0.991868 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 12016154 98 - 27905053 CAUUUAUUUUGCGUCAGCAGCGCCUCUCAAUGAGGAAACGCUGCGUAUACAUAAUUUUCUGGCCACAUAUA--CGCACUCCCAUAUCUUUCUCCCAUCGA .........(((((..((((((((((.....))))...))))))((...((........))...))....)--))))....................... ( -21.20) >DroSec_CAF1 121011 100 - 1 CAUUUAUUUUGCGUCAGCAGCGCCUCUCAAUGAGGAAACGCUGCGUAUACACAAUUUUCUGGCCACAUAUACACGCACUCCCAUAUCUUUCUCCCAUCGA .........(((((..((((((((((.....))))...))))))(((((..((......))......))))))))))....................... ( -23.10) >DroSim_CAF1 121293 100 - 1 CAUUUAUUUUGCGUCAGCAGCGCCUCUCAAUGAGGAAACGCUGCGUAUACAUAAUUUUCUGGCCACAUAUACACGCACUCCCAUAUCUUUCUCCCAUCGA .........(((((..((((((((((.....))))...))))))(((((..........((....))))))))))))....................... ( -22.80) >DroEre_CAF1 120720 98 - 1 CAUUUAUUUUGCGUCAGCAGCUCCUCACAAUGAGGAAACGCUGCGUAUACAUAAUUUUCUGGCCACAUAUA--CGCACUCUCAUAUCUUUCGGACGUGGA .......((..((((.((((((((((.....)))))...)))))(((((..........((....))))))--)..................))))..)) ( -25.50) >DroYak_CAF1 123834 98 - 1 CAUUUAUUUUGCGUCAGCAGCGCCUCUCAAUGAGGAAACGCUGCGUAUACAUAAUUUUCUGGCCACAUAUA--CGCACUCGCAUAUCUUUCGAACGUCGA .........((((...((((((((((.....))))...))))))(((((..........((....))))))--).....))))......(((.....))) ( -22.50) >consensus CAUUUAUUUUGCGUCAGCAGCGCCUCUCAAUGAGGAAACGCUGCGUAUACAUAAUUUUCUGGCCACAUAUA__CGCACUCCCAUAUCUUUCUCCCAUCGA .........((((...((((((((((.....))))...))))))((...((........))...)).......))))....................... (-19.16 = -19.36 + 0.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:31:23 2006