
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,752,049 – 11,752,149 |
| Length | 100 |
| Max. P | 0.925541 |

| Location | 11,752,049 – 11,752,149 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 72.40 |
| Mean single sequence MFE | -26.25 |
| Consensus MFE | -15.72 |
| Energy contribution | -16.78 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.603698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11752049 100 + 27905053 AUGGGAAAUAGUAACUGC--UUAAACU--CAUCGACUAUGUGAUUUUAGUCAAACGCAGAUGCCUUCAGCCACUGGAUGUGGGCAAAGGCAAGGCCUAUCUGCG (((((....(((....))--)....))--))).(((((........)))))...((((((((((((...((((.....))))((....)))))))..))))))) ( -31.20) >DroSec_CAF1 84803 95 + 1 -----AAGAAGUAACUUC--AUAAACU--CAUGGACUAUGUGAUUUUAGUCAAACGCAGAUGCCUCCAGCCACUGGAUGUGGGCAAAGGCAAGGCCUAUCUGCG -----..((((...))))--.......--....(((((........)))))...((((((((((..((.((...)).))..)))..((((...))))))))))) ( -28.80) >DroSim_CAF1 86501 95 + 1 -----AAAAAGUAACUUC--AUAAACU--CAUCGACUAUGUGAUUUUAGUCAAACGCAGAUGCCUCCAGCCACUGGAUGUGGGCAAAGGCAAGGCCUAUCUGCG -----.............--.......--....(((((........)))))...((((((((((..((.((...)).))..)))..((((...))))))))))) ( -27.50) >DroEre_CAF1 85162 83 + 1 ----------------A---CCUAACU--AAUCGACCGUGCGAUUUUAGCCAAACGCAGAUGCCUUAGGCCCCUGCAUGUGGGCAAAGGUAAGGCCUAUAUGCG ----------------.---(((((..--.(((...(((((.......))...)))..)))...))))).....((((((((((.........)))))))))). ( -24.60) >DroYak_CAF1 88869 99 + 1 UUGUGAAGAAGUAACUA---UUAAACU--CAUCUGCCAUGCGAUUUAAGUCAAACCCAGAUGCCUUAAGCCACUGGAUGUGGGCAAAGGUAAGGCGUAUCUGCG ..(((.((((((.....---....)))--..)))..)))(((((....)))........((((((((.(((((.....)).))).....))))))))....)). ( -19.00) >DroPer_CAF1 91096 99 + 1 -----GGAGAGCAAACCCACAGAAACUUUUAAUAAAUAUUGGUUUUUAGUUAAGAAGAGAUGCCUGCUGCCGCUGGAACAGGGCAACGGCAAGGCCAUCCUCCG -----((((...........(((((((..((.....))..)))))))...........(((((((((((((.(((...))))))...))).))).)))))))). ( -26.40) >consensus _____AAAAAGUAACUAC__AUAAACU__CAUCGACUAUGUGAUUUUAGUCAAACGCAGAUGCCUUCAGCCACUGGAUGUGGGCAAAGGCAAGGCCUAUCUGCG .................................(((((........)))))...(((((((((((...(((..((........))..))).))))..))))))) (-15.72 = -16.78 + 1.06)



| Location | 11,752,049 – 11,752,149 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 72.40 |
| Mean single sequence MFE | -26.35 |
| Consensus MFE | -18.14 |
| Energy contribution | -18.95 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925541 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11752049 100 - 27905053 CGCAGAUAGGCCUUGCCUUUGCCCACAUCCAGUGGCUGAAGGCAUCUGCGUUUGACUAAAAUCACAUAGUCGAUG--AGUUUAA--GCAGUUACUAUUUCCCAU .(.((((((....(((((((..((((.....))))..))))))).((((....((((...(((........))).--))))...--))))...)))))).)... ( -29.00) >DroSec_CAF1 84803 95 - 1 CGCAGAUAGGCCUUGCCUUUGCCCACAUCCAGUGGCUGGAGGCAUCUGCGUUUGACUAAAAUCACAUAGUCCAUG--AGUUUAU--GAAGUUACUUCUU----- ((((((.......(((((((..((((.....))))..)))))))))))))...(((((........)))))....--.......--((((...))))..----- ( -26.71) >DroSim_CAF1 86501 95 - 1 CGCAGAUAGGCCUUGCCUUUGCCCACAUCCAGUGGCUGGAGGCAUCUGCGUUUGACUAAAAUCACAUAGUCGAUG--AGUUUAU--GAAGUUACUUUUU----- ((((((.......(((((((..((((.....))))..))))))))))))).(((((((........)))))))..--.......--.............----- ( -28.01) >DroEre_CAF1 85162 83 - 1 CGCAUAUAGGCCUUACCUUUGCCCACAUGCAGGGGCCUAAGGCAUCUGCGUUUGGCUAAAAUCGCACGGUCGAUU--AGUUAGG---U---------------- ((((.....(((((((((((((......)))))))..))))))...))))((((((((..((((......)))))--)))))))---.---------------- ( -26.70) >DroYak_CAF1 88869 99 - 1 CGCAGAUACGCCUUACCUUUGCCCACAUCCAGUGGCUUAAGGCAUCUGGGUUUGACUUAAAUCGCAUGGCAGAUG--AGUUUAA---UAGUUACUUCUUCACAA ..(((((..((((((.....(((.((.....))))).)))))))))))((..((((((((((..(((.....)))--.))))).---.)))))..))....... ( -20.50) >DroPer_CAF1 91096 99 - 1 CGGAGGAUGGCCUUGCCGUUGCCCUGUUCCAGCGGCAGCAGGCAUCUCUUCUUAACUAAAAACCAAUAUUUAUUAAAAGUUUCUGUGGGUUUGCUCUCC----- .((((((((.(((((((((((........))))))))..))))))).............(((((.(((...((.....))...))).)))))...))))----- ( -27.20) >consensus CGCAGAUAGGCCUUGCCUUUGCCCACAUCCAGUGGCUGAAGGCAUCUGCGUUUGACUAAAAUCACAUAGUCGAUG__AGUUUAU__GAAGUUACUUUUU_____ ((((((...((((((((.(((........))).)))..))))).)))))).(((((((........)))))))............................... (-18.14 = -18.95 + 0.81)

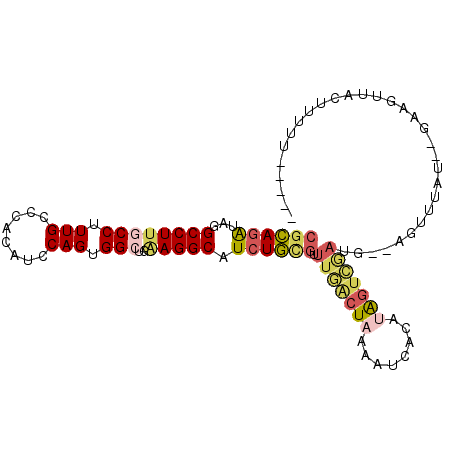

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:29:28 2006