
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,601,803 – 11,601,957 |
| Length | 154 |
| Max. P | 0.879388 |

| Location | 11,601,803 – 11,601,917 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 75.19 |
| Mean single sequence MFE | -47.14 |
| Consensus MFE | -23.12 |
| Energy contribution | -22.96 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.41 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.49 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.879388 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
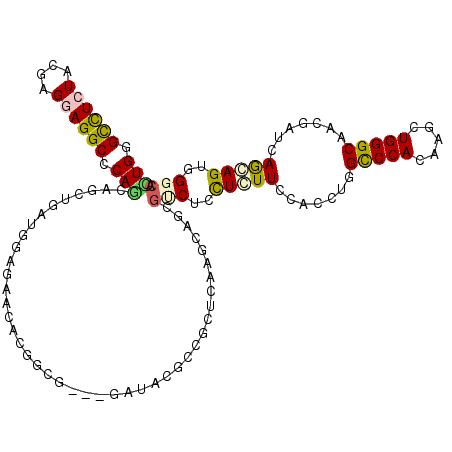
>3R_DroMel_CAF1 11601803 114 + 27905053 UUGGGCUUGUACGAGGAGGCCCAACAACUGAUGGCCAAUGCGGCU---GACAAUCCGCUGAAGCAGCGACUCCUCUUCCACUUGGCCCACAAGCUGGGCAACGAGGAGGAGUGGAGC ((((((((.(....).))))))))....((.(((((.....))))---).)).(((((((...)))).((((((((((......(((((.....)))))...))))))))))))).. ( -48.20) >DroVir_CAF1 141766 114 + 1 UUGGGCCUCUAUGAGGAGGCGCAACAGCUAAUGGAGGGCACUGCG---AAUACGCCACUGAAGCAGCGUCUCCUCUUUCAUCUCGCCCACAAGCUGGGCGAUGAUCAGCAGUGGGCG ..(((.......((((((((((....(((.......)))...(((---....)))..........))))))))))......)))((((((..(((((.......))))).)))))). ( -44.32) >DroPse_CAF1 152266 117 + 1 CUGGGUCUCUACGAGGAGGCCCAUCAACUGGUGUCGAACACAUCCGAGGAGAGUCCUCUCAAGCAGCGUCUGCUCUUCCAUUUGGUCCACAAAUUGGGCAGCGUUGAGGAGUGGGCG .(((((((((....)))))))))((..(.(((((.....))))).)..))..((((.(((...(((((.((((((....(((((.....))))).)))))))))))..))).)))). ( -49.50) >DroGri_CAF1 292685 114 + 1 CUGGGUCUCUAUGAGGAGGCACAGCAGCUGAUGGAGGACACGCCG---AAUACGCCACUCAAACAGCGUCUAUUGUUCCAUCUGGCCCACAAGCUGGGCGAUGAUCAGCAGUGGGCA (((.((((((....)))))).)))..((((...(((....((...---....))...)))...))))((((((((((.((((..(((((.....)))))))))...)))))))))). ( -41.20) >DroMoj_CAF1 135224 114 + 1 UUGGGCUUGUACGAGGAGGCACAGCAGCUGAUGGAAGGCACUGCG---GAUACGCCGCUGAAGCAGCGCCUCCUCUUUCACCUCGCCCACAAGCUGGGCGAUGAUCAACAGUGGGCA ....(((..(..((((((((.(.((.(((.......)))...(((---(.....))))....)).).))))))))..(((..(((((((.....))))))))))......)..))). ( -46.90) >DroAna_CAF1 217594 114 + 1 CUGGGCCUCUACGAGGAGGCCCAGCAGCUGAUGGAGAAGGCCGCG---GACAGCCUGCUCAAGCAGCGCCUCCUGUUUCACUUGGCCCACAAAUUGGGCAACGAAGAGCAGUGGACU ((((((((((....))))))))))..((((...(((.((((....---....)))).)))...))))..(..((((((......(((((.....)))))......))))))..)... ( -52.70) >consensus CUGGGCCUCUACGAGGAGGCCCAGCAGCUGAUGGAGAACACGGCG___GAUACGCCGCUCAAGCAGCGUCUCCUCUUCCACCUGGCCCACAAGCUGGGCAACGAUCAGCAGUGGGCA (((.((((((....)))))).)))...........................................(((..(((((.......(((((.....))))).......)))))..))). (-23.12 = -22.96 + -0.16)


| Location | 11,601,843 – 11,601,957 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 74.90 |
| Mean single sequence MFE | -47.03 |
| Consensus MFE | -22.01 |
| Energy contribution | -21.82 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.47 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.692551 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11601843 114 + 27905053 CGGCU---GACAAUCCGCUGAAGCAGCGACUCCUCUUCCACUUGGCCCACAAGCUGGGCAACGAGGAGGAGUGGAGCCAGCUGCAGGAGGAGCUGCAGGAUUCCUCGAGCUUGGAAC .((((---.......(((((...)))))((((((((((......(((((.....)))))...))))))))))..))))..((((((......))))))..((((........)))). ( -49.20) >DroVir_CAF1 141806 114 + 1 CUGCG---AAUACGCCACUGAAGCAGCGUCUCCUCUUUCAUCUCGCCCACAAGCUGGGCGAUGAUCAGCAGUGGGCGGAACUGUUGGAUGUGCUGCUCAGCACACCCAGCGUGGAGC ((((.---..............))))...((((.(.....((.(((((((..(((((.......))))).)))))))))...(((((.((((((....)))))).)))))).)))). ( -46.26) >DroPse_CAF1 152306 117 + 1 CAUCCGAGGAGAGUCCUCUCAAGCAGCGUCUGCUCUUCCAUUUGGUCCACAAAUUGGGCAGCGUUGAGGAGUGGGCGGAACUGCACGAUGAUCUGGAGAAUUCCUCGAGUGUGGAGC ((..(((((((.((((.(((...(((((.((((((....(((((.....))))).)))))))))))..))).))))....((.((.(.....))).))..)))))))..))...... ( -44.90) >DroYak_CAF1 133009 114 + 1 CGGCU---GACAACCCGCUGAAGCAGCGACUGCUCUUCCACUUGGCCCACAAGCUGGGCAGCGAGGAGGAGUGGAGCCAGCUGCAGGAGGAGCUGCAGGACUCCUCGAGCUUGGAGC .((((---.......(((((...))))).(..(((((((.(...(((((.....)))))...).)))))))..)))))..((((((......))))))..((((........)))). ( -51.20) >DroAna_CAF1 217634 114 + 1 CCGCG---GACAGCCUGCUCAAGCAGCGCCUCCUGUUUCACUUGGCCCACAAAUUGGGCAACGAAGAGCAGUGGACUGAGCUCCAAGAGGAGCUCCAGAACUCGUCCAGCUUGGAGC (((((---(((.((((((....)))).))(..((((((......(((((.....)))))......))))))..).(((((((((....)))))).))).....)))).))..))... ( -45.70) >DroPer_CAF1 154274 117 + 1 CAUCCGAGGAGAGUCCUCUCAAGCAGCGUCUGCUCUUCCAUUUGGUCCACAAAUUGGGCAGCGUUGAGGAGUGGGCGGAACUGCACGAUGAUCUGGAGAAUUCCUCGAGUGUGGAGC ((..(((((((.((((.(((...(((((.((((((....(((((.....))))).)))))))))))..))).))))....((.((.(.....))).))..)))))))..))...... ( -44.90) >consensus CAGCC___GACAGUCCGCUCAAGCAGCGUCUCCUCUUCCACUUGGCCCACAAACUGGGCAACGAUGAGGAGUGGACGGAACUGCAGGAGGAGCUGCAGAACUCCUCGAGCGUGGAGC ..............((......((((...(..((((((......(((((.....)))))......))))))..)......))))..((.(((........))).))......))... (-22.01 = -21.82 + -0.19)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:27:54 2006