
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,601,438 – 11,601,530 |
| Length | 92 |
| Max. P | 0.963334 |

| Location | 11,601,438 – 11,601,530 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 78.75 |
| Mean single sequence MFE | -17.46 |
| Consensus MFE | -9.58 |
| Energy contribution | -10.34 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.654069 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11601438 92 + 27905053 AUGCUUUUA---UUAGUCACUAUUAAAGCAGUGACGUUAAAUGUCGUGAUUUUUAACAAACAUGUGUAACUCUCAAUAAUGGUAUUAAAAGGAAA (((((.(((---((.((((((........))))))((((.((((.((........))..))))...))))....))))).))))).......... ( -16.60) >DroSec_CAF1 131243 92 + 1 AUGCUUUUA---UUAGUCACUAUUAAAGCAGUGACGUUAAAUGUCGUGAUUUUUAUCAAACUCAUGUAUCUCUUAAUACUGGUAUUAAAAGUAAA .((((((((---.((((((((........))))))(((((..(.(((((.(((....))).)))))...)..))))).....)).)))))))).. ( -18.40) >DroSim_CAF1 139421 87 + 1 UUGCUUUUA---UUAGUCACUAUUAAAGCAGUGACGUUAAAU-----GAUUUUUAUCAAACUGAUGUAUCUCUUAAUACUGGUAUUAAAAGUAAA (((((((((---.((((((((........))))))(((((..-----(((...((((.....)))).)))..))))).....)).))))))))). ( -19.90) >DroEre_CAF1 140308 72 + 1 GUGCUUUUA---UUAGUCACUAUUAAAGCAGUGAA-------------------AUC-AACAUGUGUACCUCCUAAUACUGGUACUACAGCUAAA (((((..((---(((((((((........))))).-------------------...-.((....)).....))))))..))))).......... ( -13.30) >DroYak_CAF1 132618 95 + 1 GUGCUUUUAUUAUUAGUCACUAUUAAACCAGUGACGCUAAAUGUCGUGAUUUUUAUCAAACAUGUGCACCUCUAAAUACUGGUAUUAAGACUAAA (((((..((((....((((((........))))))((...((((..((((....)))).))))..)).......))))..))))).......... ( -19.10) >consensus AUGCUUUUA___UUAGUCACUAUUAAAGCAGUGACGUUAAAUGUCGUGAUUUUUAUCAAACAUGUGUACCUCUUAAUACUGGUAUUAAAAGUAAA .((((((((......((((((........))))))..............................(((((..........))))))))))))).. ( -9.58 = -10.34 + 0.76)



| Location | 11,601,438 – 11,601,530 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 78.75 |
| Mean single sequence MFE | -17.70 |
| Consensus MFE | -11.22 |
| Energy contribution | -11.98 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963334 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11601438 92 - 27905053 UUUCCUUUUAAUACCAUUAUUGAGAGUUACACAUGUUUGUUAAAAAUCACGACAUUUAACGUCACUGCUUUAAUAGUGACUAA---UAAAAGCAU ....(((((((((....)))))))))......((((((((((((..........))))))(((((((......)))))))...---...)))))) ( -17.90) >DroSec_CAF1 131243 92 - 1 UUUACUUUUAAUACCAGUAUUAAGAGAUACAUGAGUUUGAUAAAAAUCACGACAUUUAACGUCACUGCUUUAAUAGUGACUAA---UAAAAGCAU ....(((((((((....)))))))))........(((((((....)))).))).......(((((((......)))))))...---......... ( -17.90) >DroSim_CAF1 139421 87 - 1 UUUACUUUUAAUACCAGUAUUAAGAGAUACAUCAGUUUGAUAAAAAUC-----AUUUAACGUCACUGCUUUAAUAGUGACUAA---UAAAAGCAA ....(((((((((....)))))))))........(((((((....)))-----.......(((((((......)))))))...---...)))).. ( -17.10) >DroEre_CAF1 140308 72 - 1 UUUAGCUGUAGUACCAGUAUUAGGAGGUACACAUGUU-GAU-------------------UUCACUGCUUUAAUAGUGACUAA---UAAAAGCAC .((((((((.(((((..........)))))))).)))-)).-------------------.((((((......))))))....---......... ( -15.30) >DroYak_CAF1 132618 95 - 1 UUUAGUCUUAAUACCAGUAUUUAGAGGUGCACAUGUUUGAUAAAAAUCACGACAUUUAGCGUCACUGGUUUAAUAGUGACUAAUAAUAAAAGCAC .....(((.((((....)))).))).((((..(((((((((....)))).))))).....(((((((......)))))))...........)))) ( -20.30) >consensus UUUACUUUUAAUACCAGUAUUAAGAGAUACACAUGUUUGAUAAAAAUCACGACAUUUAACGUCACUGCUUUAAUAGUGACUAA___UAAAAGCAC ....(((((((((....)))))))))..................................(((((((......)))))))............... (-11.22 = -11.98 + 0.76)

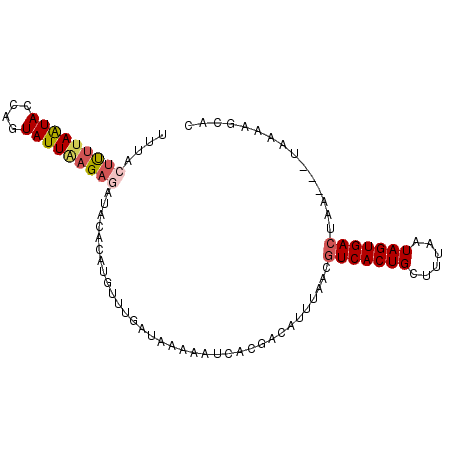

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:27:52 2006