
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,487,138 – 11,487,232 |
| Length | 94 |
| Max. P | 0.983553 |

| Location | 11,487,138 – 11,487,232 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 73.96 |
| Mean single sequence MFE | -34.80 |
| Consensus MFE | -15.44 |
| Energy contribution | -15.73 |
| Covariance contribution | 0.29 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.44 |
| SVM decision value | 1.46 |
| SVM RNA-class probability | 0.955740 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11487138 94 + 27905053 UACGGCCAACUGCAGUGUGGAAUGUGGGCUAAAAGAGGCGAACGCCU-ACCC----AUGGCAGG--GAAAUGGCC-AAGGUGGGCG--UAGUUGAGAGUUGGCU ...((((((((.(((............(((......)))..((((((-(((.----.((((...--......)))-).))))))))--)..)))..)))))))) ( -36.30) >DroSec_CAF1 21351 88 + 1 UACGGCCAACUGCAGUGUGGAAUGUGGGCUAAAA--------CGCCU-ACCC----AUGGCAGGGCGAAAUGGCC-AAGGUGGGCG--UAGUUUAGAGUUGGCU ...(((((((((((........)))...((((((--------(((((-(((.----.((((...........)))-).))))))))--)..))))))))))))) ( -35.70) >DroSim_CAF1 21461 96 + 1 UACGGCCAACUGCAGUGUGGAAUGUGGGCUAAAACAGGCCAACGCCU-ACCC----AUGGCAGGGGGAAAUGGCC-AAGGUGGGCG--UAGUUGAGAGUUGGCU ...((((((((.(((..(((..(((........)))..)))((((((-(((.----.((((...........)))-).))))))))--)..)))..)))))))) ( -40.30) >DroEre_CAF1 24671 72 + 1 UACGGCCAACUGCAGUGUGGAAUGUGGGCCAAAACCGGC-----------------------------AAUGGCC-AAGGUGGGCG--UAGUUGAGAGUUGGCU ...(((((((((((........)))..(((...((((((-----------------------------....)))-..))).))).--........)))))))) ( -24.80) >DroYak_CAF1 18635 94 + 1 UACGGCCAACUGCAGUGUGGAAUGUGGGCCAAAACGGGCGAACGCCU-ACCC----AUGGCGGG--GAAAUGGCC-AAGGUGGACG--UAGUUGAGAGCUGGCU ..(((((((((((.............(((((....((((....))))-.(((----.....)))--....)))))-..(.....))--))))))...))))... ( -32.20) >DroAna_CAF1 109053 92 + 1 UACGGCCAGCUGCAGCGUGGAAUGUAGGCCAAAAAGGAGCACUCCCGAACCCCCAGAUGGUUGG---------CUGGAGGUUGGAGGCUGGC---UGGCUGGCU ..((((((((..((((..(((.(((...((.....)).))).)))(.((((.((((........---------)))).)))).)..))))))---))))))... ( -39.50) >consensus UACGGCCAACUGCAGUGUGGAAUGUGGGCCAAAACAGGC_AACGCCU_ACCC____AUGGCAGG__GAAAUGGCC_AAGGUGGGCG__UAGUUGAGAGUUGGCU ...(((((((((((........))).(((((............(((............))).........))))).....................)))))))) (-15.44 = -15.73 + 0.29)



| Location | 11,487,138 – 11,487,232 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 73.96 |
| Mean single sequence MFE | -28.55 |
| Consensus MFE | -12.36 |
| Energy contribution | -13.67 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.43 |
| SVM decision value | 1.95 |
| SVM RNA-class probability | 0.983553 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11487138 94 - 27905053 AGCCAACUCUCAACUA--CGCCCACCUU-GGCCAUUUC--CCUGCCAU----GGGU-AGGCGUUCGCCUCUUUUAGCCCACAUUCCACACUGCAGUUGGCCGUA .(((((((.......(--((((.(((((-(((......--...)))).----))))-.)))))..((........))................))))))).... ( -27.80) >DroSec_CAF1 21351 88 - 1 AGCCAACUCUAAACUA--CGCCCACCUU-GGCCAUUUCGCCCUGCCAU----GGGU-AGGCG--------UUUUAGCCCACAUUCCACACUGCAGUUGGCCGUA .((((((((((((..(--((((.(((((-(((...........)))).----))))-.))))--------)))))).................))))))).... ( -28.02) >DroSim_CAF1 21461 96 - 1 AGCCAACUCUCAACUA--CGCCCACCUU-GGCCAUUUCCCCCUGCCAU----GGGU-AGGCGUUGGCCUGUUUUAGCCCACAUUCCACACUGCAGUUGGCCGUA .(((((((.......(--((((.(((((-(((...........)))).----))))-.))))).(((........)))...............))))))).... ( -30.60) >DroEre_CAF1 24671 72 - 1 AGCCAACUCUCAACUA--CGCCCACCUU-GGCCAUU-----------------------------GCCGGUUUUGGCCCACAUUCCACACUGCAGUUGGCCGUA .((((((.........--.(((......-)))...(-----------------------------((.(((..(((........))).)))))))))))).... ( -17.40) >DroYak_CAF1 18635 94 - 1 AGCCAGCUCUCAACUA--CGUCCACCUU-GGCCAUUUC--CCCGCCAU----GGGU-AGGCGUUCGCCCGUUUUGGCCCACAUUCCACACUGCAGUUGGCCGUA .(((((((.......(--((((.(((((-(((......--...)))).----))))-.)))))..(((......)))................))))))).... ( -28.50) >DroAna_CAF1 109053 92 - 1 AGCCAGCCA---GCCAGCCUCCAACCUCCAG---------CCAACCAUCUGGGGGUUCGGGAGUGCUCCUUUUUGGCCUACAUUCCACGCUGCAGCUGGCCGUA .((((((((---((.(((((((.((((((((---------........))))))))...)))).)))......(((........))).))))..)))))).... ( -39.00) >consensus AGCCAACUCUCAACUA__CGCCCACCUU_GGCCAUUUC__CCUGCCAU____GGGU_AGGCGUU_GCCCGUUUUAGCCCACAUUCCACACUGCAGUUGGCCGUA .(((((((...........(((.......)))....................((((..(((....))).......))))..............))))))).... (-12.36 = -13.67 + 1.31)


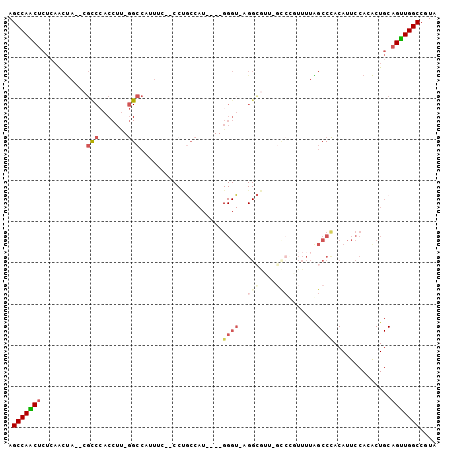
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:27:15 2006