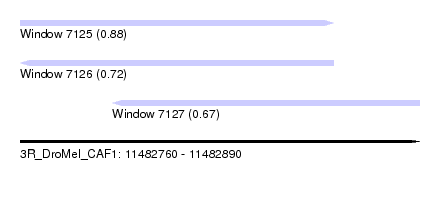
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,482,760 – 11,482,890 |
| Length | 130 |
| Max. P | 0.880487 |

| Location | 11,482,760 – 11,482,862 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 81.61 |
| Mean single sequence MFE | -23.32 |
| Consensus MFE | -20.03 |
| Energy contribution | -19.70 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.880487 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11482760 102 + 27905053 GAGCAAAAUGGGGUGAAAGUGAUUGGAUUAAAGGGAAAUCCUGGCUGAUAUCGGUGGAUAUUUCACAGGAAUUCCACGAUUACAUAAGAAGAAGGUGAUGCA ..(((..((.........(((((((........((((.(((((...((((((....))))))...))))).)))).)))))))...........))..))). ( -24.35) >DroEre_CAF1 20470 90 + 1 GAGCAAAAUGG-GUGAGAGCGAUUGGAUGAAUGGGAAAUCCUGGGUGUUAUCGCUGGAUAUUUCACAGGAAUUCCACGAUUACGCAGGCAG----------- ..((....((.-((((...(((((.....))).((((.(((((((...((((....))))..)).))))).)))).)).)))).)).))..----------- ( -23.70) >DroYak_CAF1 14342 95 + 1 GAGCGAAAUGGAGUGAAAGUGAUUGGAUGAAUGGGAAAUCCUGGGUGUUAACGCUAGAUAUUUCACAGGAAUUCCACGAUUAUGUAAGCAGAAGG------- ............((....(((((((........((((.(((((((((....)))).((....)).))))).)))).)))))))....))......------- ( -21.90) >consensus GAGCAAAAUGG_GUGAAAGUGAUUGGAUGAAUGGGAAAUCCUGGGUGUUAUCGCUGGAUAUUUCACAGGAAUUCCACGAUUACGUAAGCAGAAGG_______ ..................(((((((........((((.(((((((((....)))).((....)).))))).)))).)))))))................... (-20.03 = -19.70 + -0.33)



| Location | 11,482,760 – 11,482,862 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 81.61 |
| Mean single sequence MFE | -14.67 |
| Consensus MFE | -12.20 |
| Energy contribution | -12.53 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724127 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11482760 102 - 27905053 UGCAUCACCUUCUUCUUAUGUAAUCGUGGAAUUCCUGUGAAAUAUCCACCGAUAUCAGCCAGGAUUUCCCUUUAAUCCAAUCACUUUCACCCCAUUUUGCUC .(((.............(((....)))((((.(((((.(..(((((....)))))...)))))).))))............................))).. ( -16.40) >DroEre_CAF1 20470 90 - 1 -----------CUGCCUGCGUAAUCGUGGAAUUCCUGUGAAAUAUCCAGCGAUAACACCCAGGAUUUCCCAUUCAUCCAAUCGCUCUCAC-CCAUUUUGCUC -----------..((..(((....)))((((.(((((.(...((((....))))...).))))).)))).............))......-........... ( -15.30) >DroYak_CAF1 14342 95 - 1 -------CCUUCUGCUUACAUAAUCGUGGAAUUCCUGUGAAAUAUCUAGCGUUAACACCCAGGAUUUCCCAUUCAUCCAAUCACUUUCACUCCAUUUCGCUC -------..................(((((((....((((((........(....).....((((.........))))......))))))...))))))).. ( -12.30) >consensus _______CCUUCUGCUUACGUAAUCGUGGAAUUCCUGUGAAAUAUCCAGCGAUAACACCCAGGAUUUCCCAUUCAUCCAAUCACUUUCAC_CCAUUUUGCUC .........................(.((((.(((((.....((((....)))).....))))).)))))................................ (-12.20 = -12.53 + 0.33)



| Location | 11,482,790 – 11,482,890 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 72.07 |
| Mean single sequence MFE | -14.90 |
| Consensus MFE | -9.77 |
| Energy contribution | -10.53 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.669658 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11482790 100 - 27905053 UUA------------CUUACAAUACUAUUCAUGUUUCUACUGCAUCACCUUCUUCUUAUGUAAUCGUGGAAUUCCUGUGAAAUAUCCACCGAUAUCAGCCAGGAUUUCCCUU (((------------(..............((((.......))))..............))))..(.((((.(((((.(..(((((....)))))...)))))).))))).. ( -16.59) >DroSec_CAF1 17034 100 - 1 UUA------------CUUACAAUACUAUCCAGUUUUCUACUGCAUCACCUUCUUCUUACGUAAUCGUGGAAUUCCUGUGAAAUAUCCACCUAUAACUGCCAUGAUUUCCCUU ...------------..............((((.....)))).................(.((((((((.......(((.......))).........)))))))).).... ( -11.89) >DroEre_CAF1 20499 100 - 1 UCAGCCCCCAUUAUGCAUACAAUAAUAUCCAGACUUCUA------------CUGCCUGCGUAAUCGUGGAAUUCCUGUGAAAUAUCCAGCGAUAACACCCAGGAUUUCCCAU .........((((((((..((.((.............))------------.))..)))))))).(.((((.(((((.(...((((....))))...).))))).))))).. ( -19.02) >DroYak_CAF1 14372 99 - 1 CCA--CCCUAUUAUACAUACAAU---AGCCAGCUUUCCA--------CCUUCUGCUUACAUAAUCGUGGAAUUCCUGUGAAAUAUCUAGCGUUAACACCCAGGAUUUCCCAU ...--..(((((........)))---))..(((......--------......))).........(.((((.(((((.(...................)))))).))))).. ( -12.11) >consensus UCA____________CAUACAAUACUAUCCAGAUUUCUA________CCUUCUGCUUACGUAAUCGUGGAAUUCCUGUGAAAUAUCCACCGAUAACACCCAGGAUUUCCCAU .................................................................(.((((.(((((.....((((....)))).....))))).))))).. ( -9.77 = -10.53 + 0.75)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:27:12 2006