
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,409,974 – 11,410,165 |
| Length | 191 |
| Max. P | 0.870289 |

| Location | 11,409,974 – 11,410,087 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.20 |
| Mean single sequence MFE | -42.80 |
| Consensus MFE | -36.34 |
| Energy contribution | -36.58 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.582470 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11409974 113 + 27905053 CGGCGGUCCCGUGGGUAAAUUGAUAUUCCUGUCAUUGCUGUAAUGAUUUCUGCUUCCUGCGAUAGGAAUCGCGGUCCGGCGAUUGCCGUGUGC-------CGUGUAUCCUGGCGCAUCCU (((((((((((.((((((..(((((....)))))))))).................(((((((....))))))).)))).))))))))(((((-------((.......))))))).... ( -39.20) >DroSec_CAF1 157620 113 + 1 CGGCGGUCCCGUGGGUAAAUUGAUAUUCCUGUCAUUGCUGUAAUGAUUUCUGCUUCCUGCGGCAGGAAUCGCGGUCCGGCGAUUGCCGUGUGC-------CGUGUAUCCUGGCGCUUCCU (((((((((((.((((((..(((((....)))))))))).....((((((((((......)))))))))).....)))).)))))))).((((-------((.......))))))..... ( -42.70) >DroSim_CAF1 149088 113 + 1 CGGCGGUCCCGUGGGUAAAUUGAUAUUCCUGUCAUUGCUGUAAUGAUUUCUGCUUCCUGCGGCAGGAAUCGCGGUCCGGCGAUUGCCGUGUGC-------CGUGUAUCCUGGCGCAUCCU (((((((((((.((((((..(((((....)))))))))).....((((((((((......)))))))))).....)))).))))))))(((((-------((.......))))))).... ( -43.90) >DroEre_CAF1 145351 113 + 1 CGGCGGUCCCGUGGGUAAAUUGAUAUUCCUGUCAUUGCUGUAAUGAUUUCUGCUUCCUGCGGCAGGAACCGCGGUCCGGCGAUUGCCGUGUGC-------CGUGUAUCCUGGCGCAUCCU .(((((...(((.(((((..(((((....))))))))))...)))....)))))...((((.(((((..(((((((((((....)))).).))-------))))..))))).)))).... ( -43.50) >DroYak_CAF1 149208 120 + 1 CGGCGGUCCCGUGGGUAAAUUGAUAUUCCUGUCAUUGCUGUAAUGAUUUCUGCUUCCUGCGGCAGGAAUCGCGGUCCGGCGAUUGCCGUGUGCCGUAUGCCGUGUAUCCUGGCGCAUCCU (((((((((((.((((((..(((((....)))))))))).....((((((((((......)))))))))).....)))).))))))))(((((((.((((...))))..))))))).... ( -44.70) >consensus CGGCGGUCCCGUGGGUAAAUUGAUAUUCCUGUCAUUGCUGUAAUGAUUUCUGCUUCCUGCGGCAGGAAUCGCGGUCCGGCGAUUGCCGUGUGC_______CGUGUAUCCUGGCGCAUCCU (((((((((((.((((((..(((((....)))))))))).....((((((((((......)))))))))).....)))).)))))))).............((((......))))..... (-36.34 = -36.58 + 0.24)


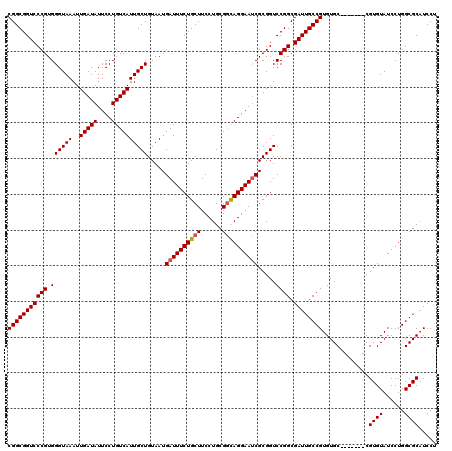
| Location | 11,410,014 – 11,410,125 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 91.30 |
| Mean single sequence MFE | -39.42 |
| Consensus MFE | -33.80 |
| Energy contribution | -33.92 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.870289 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11410014 111 - 27905053 CCUGUCGCGGAUAGAAAAAAAAACGACAAACCGCAGGCAGGAUGCGCCAGGAUACACG-------GCACACGGCAAUCGCCGGACCGCGAUUCCUAUCGCAGGAAGCAGAAAUCAUUA ((((((((((....................)))).)))))).((((((.(......))-------))...((((....))))..(((((((....))))).))..))).......... ( -36.25) >DroEre_CAF1 145391 108 - 1 CCUGUCGCGGAUAGAAA-A--AACGACCAGCCGCAGGCAGGAUGCGCCAGGAUACACG-------GCACACGGCAAUCGCCGGACCGCGGUUCCUGCCGCAGGAAGCAGAAAUCAUUA ((((((((((.......-.--.........)))).)))))).((((.(((((..(.((-------(....((((....))))..))).)..))))).))))................. ( -40.93) >DroYak_CAF1 149248 116 - 1 CCUGUCGCGGAUAGAAAAA--AACGACAAACCGCAGGCAGGAUGCGCCAGGAUACACGGCAUACGGCACACGGCAAUCGCCGGACCGCGAUUCCUGCCGCAGGAAGCAGAAAUCAUUA (((((.((((.........--.........)))).(((((((((((((.(......))))....((....((((....))))..)))))..))))))))))))............... ( -41.07) >consensus CCUGUCGCGGAUAGAAAAA__AACGACAAACCGCAGGCAGGAUGCGCCAGGAUACACG_______GCACACGGCAAUCGCCGGACCGCGAUUCCUGCCGCAGGAAGCAGAAAUCAUUA (((((.((((....................)))).(((((((..(((..(......).............((((....))))....)))..))))))))))))............... (-33.80 = -33.92 + 0.11)

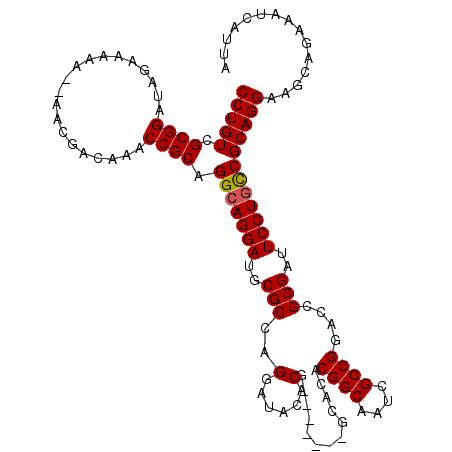
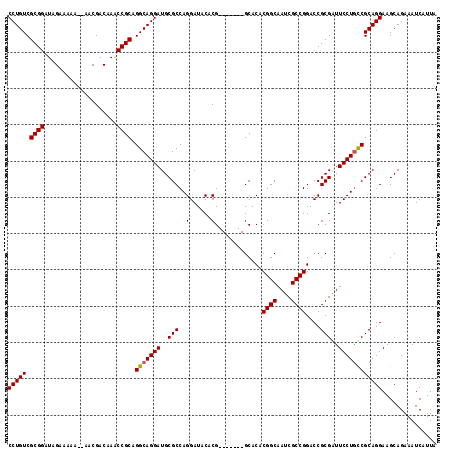
| Location | 11,410,054 – 11,410,165 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.22 |
| Mean single sequence MFE | -28.64 |
| Consensus MFE | -22.34 |
| Energy contribution | -22.74 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.593817 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11410054 111 - 27905053 UAAUCUCGAAGAAAAUGUUUGCCUUUCUUUAAUUCAAUAUCCUGUCGCGGAUAGAAAAAAA--AACGACAAACCGCAGGCAGGAUGCGCCAGGAUACACG-------GCACACGGCAAUC ..................(((((..............(((((((((((((...........--.........)))).))))))))).(((.(......))-------))....))))).. ( -29.05) >DroSec_CAF1 157700 113 - 1 UAAUCUCGAAGAAAAUGUUUGCCUUUCUUUAAUUCAAUAUCCUGUCGCGGAUAUAAAAAAAAGAACGACAAGCGGCAGGCAGGAAGCGCCAGGAUACACG-------GCACACGGCAAUC ..................(((((.........(((.....((((((((.......................))))))))...)))(.(((.(......))-------)).)..))))).. ( -25.30) >DroSim_CAF1 149168 113 - 1 UAAUCUCGAAGAAAAUGUUUGCCUUUCUUUAAUUCAAUAUCCUGUCGCGGAUAGAAAAAAAAGAACGACAAACCGCAGGCAGGAUGCGCCAGGAUACACG-------GCACACGGCAAUC ..................(((((..............(((((((((((((......................)))).))))))))).(((.(......))-------))....))))).. ( -28.95) >DroEre_CAF1 145431 108 - 1 UAAUCUCGAAGAAAAUGUUUGCCUUUCUUUAAUUCAAUAUCCUGUCGCGGAUAGAAA-A----AACGACCAGCCGCAGGCAGGAUGCGCCAGGAUACACG-------GCACACGGCAAUC ..................(((((..............(((((((((((((.......-.----.........)))).))))))))).(((.(......))-------))....))))).. ( -29.43) >DroYak_CAF1 149288 116 - 1 UAAUCUCGAAGAAAAUGUUUGCCUUUCUUUAAUUCAAUAUCCUGUCGCGGAUAGAAAAA----AACGACAAACCGCAGGCAGGAUGCGCCAGGAUACACGGCAUACGGCACACGGCAAUC ...............(((.((((..............(((((((((((((.........----.........)))).))))))))).(((.(......))))....)))).)))...... ( -30.47) >consensus UAAUCUCGAAGAAAAUGUUUGCCUUUCUUUAAUUCAAUAUCCUGUCGCGGAUAGAAAAAAA__AACGACAAACCGCAGGCAGGAUGCGCCAGGAUACACG_______GCACACGGCAAUC ..................(((((..............(((((((((((((......................)))).))))))))).((..................))....))))).. (-22.34 = -22.74 + 0.40)

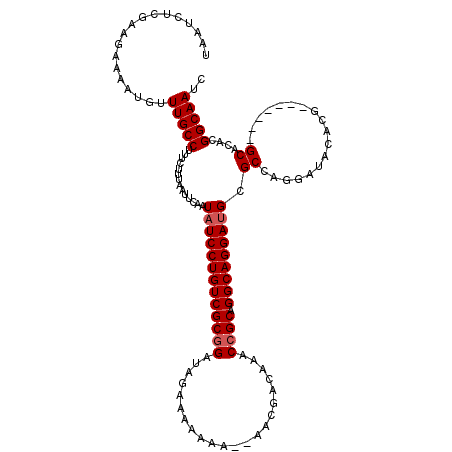
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:24:42 2006