
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,399,239 – 11,399,589 |
| Length | 350 |
| Max. P | 0.835978 |

| Location | 11,399,239 – 11,399,338 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.32 |
| Mean single sequence MFE | -27.18 |
| Consensus MFE | -24.76 |
| Energy contribution | -24.96 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.835978 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11399239 99 - 27905053 CUUCUCAGCUGAUCAAAUCACAAC---------------------GCACAUUGCACACGCUCGCCGCCGGAAUAAAACAUUUCCAUGAGCUGACGAUUUGACAGGCAUCCGGGCAGGGAA ..((((..(((.(((((((.....---------------------((.....)).((.(((((.....((((........)))).)))))))..))))))))))((......)).)))). ( -26.50) >DroSec_CAF1 146928 99 - 1 CUUCUCAGCUGAUCAAAUCACAAC---------------------GCACAUUGCACACGCUCGCCGCCGGAAUAAAACAUUUCCAUGAGCUGACGAUUUGACAGGCAUCCGGCCAGGGAA ..((((...((.(((((((.....---------------------((.....)).((.(((((.....((((........)))).)))))))..)))))))))(((.....))).)))). ( -26.60) >DroSim_CAF1 138016 99 - 1 CUUCUCAGCUGAUCAAAUCACAAC---------------------GCACAUUGCACACGCUCGCCGCCGGAAUAAAACAUUUCCAUGAGCUGACGAUUUGACAGGCAUCCGGCCAGGGAA ..((((...((.(((((((.....---------------------((.....)).((.(((((.....((((........)))).)))))))..)))))))))(((.....))).)))). ( -26.60) >DroEre_CAF1 135040 120 - 1 CUUCUCAGCUGAUCAAAUCACAACGCACAACGCACAAGGCACAACGCACAUUGCACACGCUCGCCGCCGGAAUAAAACAUUUCCAUGAGCUGACGAUUUGACAGGCAUCCGCCCAGGGAA .......(((..(((((((............(((....((.....))....))).((.(((((.....((((........)))).)))))))..)))))))..))).(((......))). ( -29.20) >DroYak_CAF1 137954 99 - 1 AUUCUCAGCUGAUCAAAUCACAAC---------------------GCACAUUGCACACGCUCGCCGCCGGAAUAAAACAUUUCCAUGAGCUGACGAUUUGACAGGCAGCCGCGCAGGGAA .(((((.(((..(((((((.....---------------------((.....)).((.(((((.....((((........)))).)))))))..)))))))..))).((...)).))))) ( -27.00) >consensus CUUCUCAGCUGAUCAAAUCACAAC_____________________GCACAUUGCACACGCUCGCCGCCGGAAUAAAACAUUUCCAUGAGCUGACGAUUUGACAGGCAUCCGGCCAGGGAA .......(((..(((((((..........................((.....)).((.(((((.....((((........)))).)))))))..)))))))..))).(((......))). (-24.76 = -24.96 + 0.20)

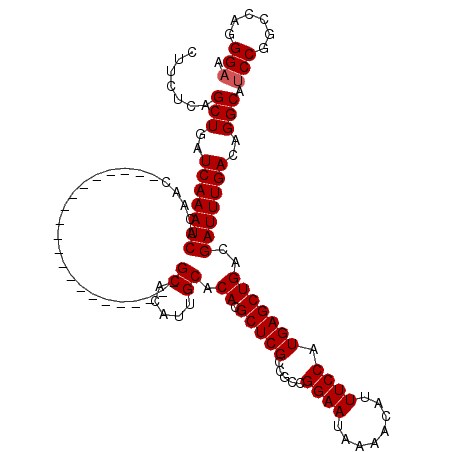

| Location | 11,399,338 – 11,399,436 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 88.43 |
| Mean single sequence MFE | -17.18 |
| Consensus MFE | -13.43 |
| Energy contribution | -13.68 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.577644 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11399338 98 - 27905053 CAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA-AGACAGAGCCACAUCCACUUCCACAUCCACAUCCAGAUGCAGAGUCAAAUCCCACCCACGCACACA ...(((((((........))))))).((((..-.....((......)).((((...((((........))))..))))..............))))... ( -19.30) >DroSec_CAF1 147027 85 - 1 CAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA-AGACAGAGCCACA------------UCCACAUCCAGAUGCAGAGUCAAAUCCCACCCACGC-CACA ...(((((((........))))))).......-.(((....(..((------------((........))))..).)))...............-.... ( -14.90) >DroSim_CAF1 138115 86 - 1 CAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA-AGACAGAGCCACA------------UCCACAUCCAGAUGCAGAGUCAAAUCCCACCCACGCACACA ...(((((((........))))))).((((..-...........((------------((........))))..((......))........))))... ( -17.70) >DroEre_CAF1 135160 99 - 1 CAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCACAGCCACAGCCACAUCCACAUCCACAUCCACAUCCAGAUCCUGAGUCAAAUCCCACCCACGCACACA ...(((((((........))))))).((((....((....))..........................(((.....))).............))))... ( -16.80) >consensus CAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA_AGACAGAGCCACA____________UCCACAUCCAGAUGCAGAGUCAAAUCCCACCCACGCACACA ...(((((((........))))))).((((....(.......).........................(((.....))).............))))... (-13.43 = -13.68 + 0.25)



| Location | 11,399,357 – 11,399,474 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.01 |
| Mean single sequence MFE | -35.70 |
| Consensus MFE | -28.08 |
| Energy contribution | -29.08 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.558557 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11399357 117 - 27905053 -UGAAUGCAGGCCAUCAAUGG-GGCCAGGACUGGAGCUGCCAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA-AGACAGAGCCACAUCCACUUCCACAUCCACAUCCAGAUGCAGAGUCA -.((.((((((((........-)))).(((.(((((.((....(((((((........))))))).(((.(.-.......).)))...)).)))))..))).........))))...)). ( -34.60) >DroSec_CAF1 147045 106 - 1 -UGGAUGCAGGCCAUCAAUGGGGGCCAGGACUGGAGCUGCCAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA-AGACAGAGCCACA------------UCCACAUCCAGAUGCAGAGUCA -((((((..((((.((....)))))).(((.(((..((((((.(((((((........)))))))..))...-.).)))..)))..------------))).))))))(((.....))). ( -36.60) >DroSim_CAF1 138134 106 - 1 -UGGAUGCAGGCCAUCAAUGGGGGCCAGGACUGGAGCUGCCAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA-AGACAGAGCCACA------------UCCACAUCCAGAUGCAGAGUCA -((((((..((((.((....)))))).(((.(((..((((((.(((((((........)))))))..))...-.).)))..)))..------------))).))))))(((.....))). ( -36.60) >DroEre_CAF1 135179 115 - 1 ----AUGCAGGCCAUCAAUGG-GGCCAGGACUGGAGCUGCCAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCACAGCCACAGCCACAUCCACAUCCACAUCCACAUCCAGAUCCUGAGUCA ----.......(((....)))-(((((((((((((..((....(((((((........))))))).(((.....((....))......))).........))..))))).))))).))). ( -33.50) >DroYak_CAF1 138093 106 - 1 GUGGAUGCACGCCAUCAAUGG-GGCCAGGACUGGAGCUGCCAGCUGCCAUAAUCAUUAGUGGCAGAGAGCCA-AGACUGAGCCAC------------AUCCACAUCCAGAUCCUGAGUCA (((((((....(((....)))-((((((...(((..((.....(((((((........)))))))))..)))-...))).))).)------------)))))).....(((.....))). ( -37.20) >consensus _UGGAUGCAGGCCAUCAAUGG_GGCCAGGACUGGAGCUGCCAGCUGCCAUAAUCAUUAGUGGCAGAGUGCCA_AGACAGAGCCACA___________AUCCACAUCCAGAUGCAGAGUCA .((((((..((((.(.....).)))).(((((((.....))))(((((((........))))))).................................))).))))))(((.....))). (-28.08 = -29.08 + 1.00)

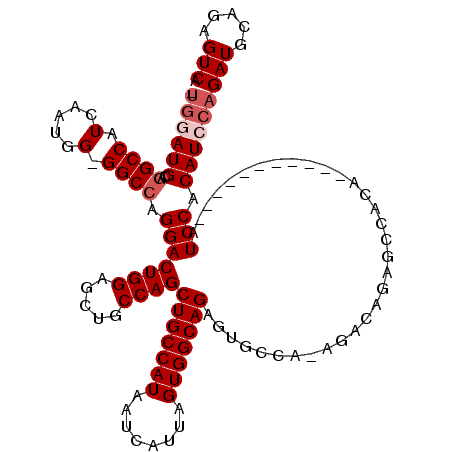

| Location | 11,399,474 – 11,399,589 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.05 |
| Mean single sequence MFE | -30.07 |
| Consensus MFE | -25.83 |
| Energy contribution | -26.70 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.557043 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11399474 115 - 27905053 UGAUUCGCCGGACUUGGCAUUUCCGCCCACCCACAAUCACAUUCAUCCAAGCUCAGUCAACACUCGAAACUCACUUGUUUCGCCAUCGUUUUGCGACUGCAGUUCGUGGUGCAUG----- (((((.(..((....(((......)))..))..))))))(((.(((((.(((((((((.......(((((......)))))((.........))))))).)))).).)))).)))----- ( -27.40) >DroSec_CAF1 147151 115 - 1 UGAUUCGCCGGACUUGGCAUUUCCGCCCACCCACAAUCACAUUCAUCCAAGCUCAGUCAACACUCGAAACUCACUUGUUUCGCCAUCGUUUUGCGACUGCAGUUCGUGGUUGAUG----- (((((.(..((....(((......)))..))..))))))...(((.(((.((((((((.......(((((......)))))((.........))))))).)))...))).)))..----- ( -27.00) >DroSim_CAF1 138240 115 - 1 UGAUUCACCGGACUUGGCAUUUCCGCCCACCCACAAUCACAUUCAUCCAAGCUCAGUCAACACUCGAAACUCACUUGUUUCGCCAUCGUUUUGCGACUGCAGUUCGUGGUGGAUG----- (((((....((....(((......)))...))..)))))(((((((((.(((((((((.......(((((......)))))((.........))))))).)))).).))))))))----- ( -30.90) >DroYak_CAF1 138199 120 - 1 UGACUCACCGGACUUGGCACUUCCGCCCACCCAAAAUCGCAUUCAUCCGAGCUCAGUAAACACUCGAAACUCACUUGUUUCGCCAUCGUUUUGCGACUGCAUCUCGUGGUGGAUGUGGAU .........((....(((......)))...))....(((((((((((((((..((((.......((((((......))))))....((.....))))))...)))).))))))))))).. ( -35.00) >consensus UGAUUCACCGGACUUGGCAUUUCCGCCCACCCACAAUCACAUUCAUCCAAGCUCAGUCAACACUCGAAACUCACUUGUUUCGCCAUCGUUUUGCGACUGCAGUUCGUGGUGGAUG_____ (((((....((....(((......)))...))..)))))(((((((((.(((((((((.......(((((......)))))((.........))))))).)))).).))))))))..... (-25.83 = -26.70 + 0.87)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:24:04 2006