
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,359,964 – 11,360,324 |
| Length | 360 |
| Max. P | 0.971390 |

| Location | 11,359,964 – 11,360,084 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.25 |
| Mean single sequence MFE | -40.82 |
| Consensus MFE | -36.98 |
| Energy contribution | -37.42 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.971390 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11359964 120 + 27905053 AGGGUAGUAUUUCUUCCGCUCCGCCCUCGUAGGUCAGUUUCACCAGCUCUGCUGCCCGAUUGGCCAGCGAAUUGGAGUUGGCUAGGAUUAUGCCCACGGGCUGACCAUAAAACUUAAUCU .(((((....((((.((((((((...((((.((((((((....((((...))))...)))))))).))))..))))).)))..))))...)))))..((.....)).............. ( -40.90) >DroSec_CAF1 107512 120 + 1 UGGGUAGUAUUUCUUCCGCUCCGCCUUCGUAGGUCAGUUUCACCAGCUCUGCUGCCCGAUUGGCCAGCGAAUUGGAGUUGGCUAGGAUUAUGCCCACGGGUUGACCAUAAAACUUAAUCU ((((((....((((.((((((((..(((((.((((((((....((((...))))...)))))))).))))).))))).)))..))))...)))))).((.....)).............. ( -42.00) >DroSim_CAF1 98294 120 + 1 UGGGUAGUAUCUCUUCCGCUCCGCCCUCGUAGGUCAGUUUCACCAGCUCUGCUGCCCGAUUGGCCAGCGAAUUGGAGUUGGCUAGAAUUAUGCCCACAGGCUGACCAUAAAACUUAAUCA ((((((.((..(((.((((((((...((((.((((((((....((((...))))...)))))))).))))..))))).)))..)))..))))))))..((....)).............. ( -38.70) >DroEre_CAF1 94942 120 + 1 UGGGCAGUAUUUCUUCCGCUCCGCCCUCGUAGGUCAGUUUCACCAGCUCUGCGGCCCGAUUGGCCAGCGAAUUGGAGUUGGCUAGGAUCAUGCCCACGGGCUGACCAUAGAACAUAAUUU ((((((....((((.((((((((...((((.((((((((...((.((...))))...)))))))).))))..))))).)))..))))...)))))).((.....)).............. ( -41.50) >DroYak_CAF1 97617 120 + 1 UGGGCAGUACUUCUUCCGCUCCACCCUCGUAGCUCAGUUUCACCAGCUCUGCGGCCCGAUUGGCCAGCGAAUUGGAGUUGGCUAGGAUCAUGCCCACGGGCUGACCAUAGAACGAAAUUU ((((((....((((.((((((((...((((.((..((((.....))))..))((((.....)))).))))..))))).)))..))))...))))))((..(((....)))..))...... ( -41.00) >consensus UGGGUAGUAUUUCUUCCGCUCCGCCCUCGUAGGUCAGUUUCACCAGCUCUGCUGCCCGAUUGGCCAGCGAAUUGGAGUUGGCUAGGAUUAUGCCCACGGGCUGACCAUAAAACUUAAUCU .(((((....((((.((((((((...((((.((((((((....((((...))))...)))))))).))))..))))).)))..))))...)))))..((.....)).............. (-36.98 = -37.42 + 0.44)



| Location | 11,360,084 – 11,360,204 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.50 |
| Mean single sequence MFE | -44.74 |
| Consensus MFE | -33.90 |
| Energy contribution | -34.66 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.555187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11360084 120 - 27905053 AUCUACCCGGGGUGGUGGCGUACCUCGAUGCCAAGGACAUUCCGGGACCGAACUACGUUGGGCCCAAAAUCCGGGACCAGUUUUUCUUCCCCAAGGAUGAGGAACUAUUCGCCACGGGAG .....(((((((((.((((((......))))))....))))))))).(((..........(((((.......))).))((((((((.(((....))).))))))))........)))... ( -46.20) >DroSec_CAF1 107632 120 - 1 AUCUACCCGGGGUGGUGGCGUAUCUCGACGCCAAGGACAUUCCGGGACCCAACUACGUUGGGCCCAAAAUCCGCGACACUCAUUUCUUCCCCAAGGAUGAGGAACUAUUCGCCACAGGAG .....(((((((((.((((((......))))))....))))))))).((((((...)))))).......(((((((..((((((.(((....))))))))).......))))....))). ( -43.80) >DroSim_CAF1 98414 120 - 1 AUCUACCCGGGGUGGUGGCGUACCUCGACGCCAAGGACAUUCCGGGACCCAACUACGUUGGGCCCAAAAUCCGCGACGACUUUUUCUUCCCCAAUGAUGAGGAACUAUUCGCCACAGGAG .....(((((((((.((((((......))))))....))))))))).((((((...)))))).......(((((((....((((((.((......)).))))))....))))....))). ( -42.80) >DroEre_CAF1 95062 120 - 1 AUCUACCCGGAGUGGUGGCAUACCUCGAUGCCAAGGACAUUCCGGGACCCAAUUAUGUGGGGCCCAAAUUAAGGGACGCGCACUUCUUCCCACAGGAUGAGGAACUGUUCGCCGCAGGUC .....(((((((((.((((((......))))))....))))))))).........((((.(.(((.......))).).))))((((.(((....))).))))..((((.....))))... ( -45.30) >DroYak_CAF1 97737 120 - 1 AUCUACCCGGCGUGGUGGCGUACCUCGAUGCCAAGGACAUUCCGGGACCCAACUACGUCGGCCCCAAGGUCAGGGACGCGUUCUUCUUCCCGCAGGACGAGCAACUGUUCGCCACGGGUC .....(((((((((((((.((.(((.(((((....).)).)).))))))).))))))))((((....)))).))).............((((..((.(((((....))))))).)))).. ( -45.60) >consensus AUCUACCCGGGGUGGUGGCGUACCUCGAUGCCAAGGACAUUCCGGGACCCAACUACGUUGGGCCCAAAAUCCGGGACGCGUUUUUCUUCCCCAAGGAUGAGGAACUAUUCGCCACAGGAG .....(((((((((.((((((......))))))....))))))))).((((((...)))))).((.......((((...........))))...((.((((......))))))...)).. (-33.90 = -34.66 + 0.76)


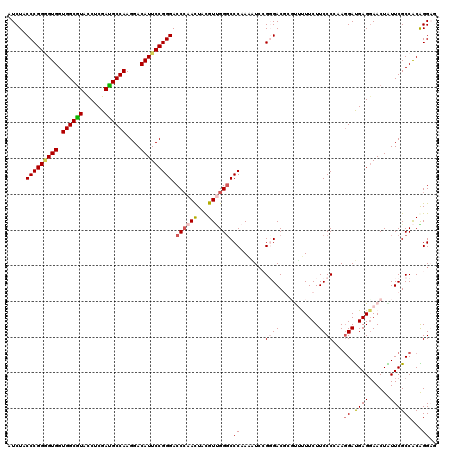
| Location | 11,360,124 – 11,360,244 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.92 |
| Mean single sequence MFE | -46.80 |
| Consensus MFE | -38.74 |
| Energy contribution | -39.22 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.798826 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11360124 120 - 27905053 GGCGCAAAAGUCACCAAGGUGGAUACGCAACCGGCUCUGGAUCUACCCGGGGUGGUGGCGUACCUCGAUGCCAAGGACAUUCCGGGACCGAACUACGUUGGGCCCAAAAUCCGGGACCAG .(((......((((....))))...)))....((..((((((...(((((((((.((((((......))))))....))))))))).((((......)))).......))))))..)).. ( -46.60) >DroSec_CAF1 107672 120 - 1 GGCGCCAAUGUAACCAAAGUGGAUACGCAACCAGCUCUGGAUCUACCCGGGGUGGUGGCGUAUCUCGACGCCAAGGACAUUCCGGGACCCAACUACGUUGGGCCCAAAAUCCGCGACACU .................(((((((((((.((((.((((((......)))))))))).)))))))....(((...((.....))(((.((((((...))))))))).......))).)))) ( -48.70) >DroSim_CAF1 98454 120 - 1 GGCGCCAAAGUCACCAAAGUGGAUACGCAACCAGCUCUGGAUCUACCCGGGGUGGUGGCGUACCUCGACGCCAAGGACAUUCCGGGACCCAACUACGUUGGGCCCAAAAUCCGCGACGAC (((......)))........((.(((((.((((.((((((......)))))))))).)))))))(((.(((...((.....))(((.((((((...))))))))).......))).))). ( -48.50) >DroEre_CAF1 95102 120 - 1 GGCGCCAAGGUCACCAAAGUGGAUACGCAGCCAGCUCUGGAUCUACCCGGAGUGGUGGCAUACCUCGAUGCCAAGGACAUUCCGGGACCCAAUUAUGUGGGGCCCAAAUUAAGGGACGCG .(((((...(((......(((....))).((((.((((((......))))))))))(((((......)))))...))).....(((.((((......)))).)))........)).))). ( -45.90) >DroYak_CAF1 97777 120 - 1 GGCGCCAAGGUCACCAAAGUGGAUACGCAGCCAGCUCUGGAUCUACCCGGCGUGGUGGCGUACCUCGAUGCCAAGGACAUUCCGGGACCCAACUACGUCGGCCCCAAGGUCAGGGACGCG .(((((..((((.((...(.((.(((((.((((.(.((((......)))).))))).))))))).)........((.....))))))))..........((((....))))..)).))). ( -44.30) >consensus GGCGCCAAAGUCACCAAAGUGGAUACGCAACCAGCUCUGGAUCUACCCGGGGUGGUGGCGUACCUCGAUGCCAAGGACAUUCCGGGACCCAACUACGUUGGGCCCAAAAUCCGGGACGCG ((((......((((....)))).(((((.((((.((((((......)))))))))).)))))......))))..((.....))(((.((((((...)))))))))............... (-38.74 = -39.22 + 0.48)



| Location | 11,360,164 – 11,360,284 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.75 |
| Mean single sequence MFE | -46.44 |
| Consensus MFE | -40.26 |
| Energy contribution | -40.02 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.683507 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11360164 120 - 27905053 CAACCAGUUGUGGGCUGCGUUUGUCAUCGCGAAGAAGGUGGGCGCAAAAGUCACCAAGGUGGAUACGCAACCGGCUCUGGAUCUACCCGGGGUGGUGGCGUACCUCGAUGCCAAGGACAU ......(((.(.(((((((((..((.((.....)).))..))))))...(((((....))((.(((((.((((.((((((......)))))))))).)))))))..)))))).).))).. ( -46.10) >DroSec_CAF1 107712 120 - 1 CAACCAGUUGUGGGCUGCAUUUGUCACCGCGAAGAAGGUGGGCGCCAAUGUAACCAAAGUGGAUACGCAACCAGCUCUGGAUCUACCCGGGGUGGUGGCGUAUCUCGACGCCAAGGACAU ......(((.(.(((((((((.((..((((.......))))..)).))))))......((((((((((.((((.((((((......)))))))))).))))))))..))))).).))).. ( -46.10) >DroSim_CAF1 98494 120 - 1 CAACCAGUUGUGGGCUGCGUUUGUCACCGCGAAGAAGGUGGGCGCCAAAGUCACCAAAGUGGAUACGCAACCAGCUCUGGAUCUACCCGGGGUGGUGGCGUACCUCGACGCCAAGGACAU ......(((.(.(((.((((...(((((........)))))))))....(((.((.....)).(((((.((((.((((((......)))))))))).)))))....)))))).).))).. ( -44.90) >DroEre_CAF1 95142 120 - 1 CAACCAGUUGUGGGCUGCGUUCGUCACCGCGAAGAAGGUGGGCGCCAAGGUCACCAAAGUGGAUACGCAGCCAGCUCUGGAUCUACCCGGAGUGGUGGCAUACCUCGAUGCCAAGGACAU ..(((.......((((((((...(((((.....)..(((((.(.....).)))))...))))..)))))))).(((((((......))))))))))(((((......)))))........ ( -47.80) >DroYak_CAF1 97817 120 - 1 CAACCAGUUGUGGGCUGCGUUUGUCACCGCGAAGAAGGUGGGCGCCAAGGUCACCAAAGUGGAUACGCAGCCAGCUCUGGAUCUACCCGGCGUGGUGGCGUACCUCGAUGCCAAGGACAU ...((((..((.((((((((...(((((.....)..(((((.(.....).)))))...))))..)))))))).)).)))).....((.(((((.(.((....)).).)))))..)).... ( -47.30) >consensus CAACCAGUUGUGGGCUGCGUUUGUCACCGCGAAGAAGGUGGGCGCCAAAGUCACCAAAGUGGAUACGCAACCAGCUCUGGAUCUACCCGGGGUGGUGGCGUACCUCGAUGCCAAGGACAU ......(((.(.(((.((((...(((((........))))))))).............(.((.(((((.((((.((((((......)))))))))).))))))).)...))).).))).. (-40.26 = -40.02 + -0.24)



| Location | 11,360,204 – 11,360,324 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.50 |
| Mean single sequence MFE | -48.92 |
| Consensus MFE | -47.18 |
| Energy contribution | -47.62 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.729437 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11360204 120 + 27905053 CCAGAGCCGGUUGCGUAUCCACCUUGGUGACUUUUGCGCCCACCUUCUUCGCGAUGACAAACGCAGCCCACAACUGGUUGUGCUGGGUGGGCAGAUCGUUGGAGUAGGUGGCUUCGCCAG ...(((((((..(((((..(((....))).....)))))))((((.(((((((........))).((((((..(..(.....)..)))))))........)))).)))))))))...... ( -43.30) >DroSec_CAF1 107752 120 + 1 CCAGAGCUGGUUGCGUAUCCACUUUGGUUACAUUGGCGCCCACCUUCUUCGCGGUGACAAAUGCAGCCCACAACUGGUUGUGCUGGGUGGGCAGAUCGUUGGAGUAGGUGGCUUCGCCAG ((((((.(((........)))))))))......(((((.((((((.(((((((....).......((((((..(..(.....)..))))))).....).))))).))))))...))))). ( -49.00) >DroSim_CAF1 98534 120 + 1 CCAGAGCUGGUUGCGUAUCCACUUUGGUGACUUUGGCGCCCACCUUCUUCGCGGUGACAAACGCAGCCCACAACUGGUUGUGCUGGGUGGGCAGAUCGUUGGAGUAGGUGGCUUCGCCAG ((((((.(((........)))))))))......(((((.((((((.(((((((........))).((((((..(..(.....)..)))))))........)))).))))))...))))). ( -51.40) >DroEre_CAF1 95182 120 + 1 CCAGAGCUGGCUGCGUAUCCACUUUGGUGACCUUGGCGCCCACCUUCUUCGCGGUGACGAACGCAGCCCACAACUGGUUGUGCUGGGUGGGCAGAUCGUUGGAGUAGGUGGCUUCGCCAG ((((((.(((........)))))))))......(((((.((((((.((((.((....))((((..((((((..(..(.....)..)))))))....)))))))).))))))...))))). ( -50.50) >DroYak_CAF1 97857 120 + 1 CCAGAGCUGGCUGCGUAUCCACUUUGGUGACCUUGGCGCCCACCUUCUUCGCGGUGACAAACGCAGCCCACAACUGGUUGUGCUGGGUGGGCAGAUCGUUGGAGUAGGUGGCUUCGCCAG ((((((.(((........)))))))))......(((((.((((((.(((((((........))).((((((..(..(.....)..)))))))........)))).))))))...))))). ( -50.40) >consensus CCAGAGCUGGUUGCGUAUCCACUUUGGUGACCUUGGCGCCCACCUUCUUCGCGGUGACAAACGCAGCCCACAACUGGUUGUGCUGGGUGGGCAGAUCGUUGGAGUAGGUGGCUUCGCCAG ((((((.(((........)))))))))......(((((.((((((.(((((((........))).((((((..(..(.....)..)))))))........)))).))))))...))))). (-47.18 = -47.62 + 0.44)


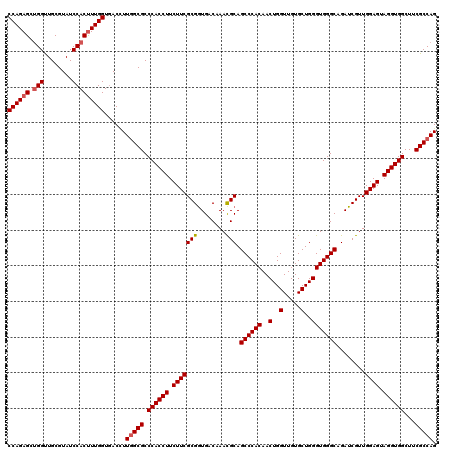
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:22:09 2006