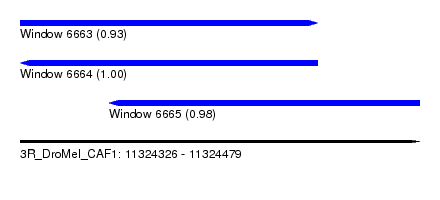

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,324,326 – 11,324,479 |
| Length | 153 |
| Max. P | 0.999967 |

| Location | 11,324,326 – 11,324,440 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.85 |
| Mean single sequence MFE | -27.90 |
| Consensus MFE | -25.66 |
| Energy contribution | -26.10 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.931222 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11324326 114 + 27905053 ------GGGGCAGGAGGAGUCCCUGGCUAAAUUUAAUUUCGCCCCAAAGAAAAAUCACACGAAAUCGACACCUGCCGUGGCGCCGGAAAAUAAUCAAAAUAAAAUAUGGAUUUUUCUUUG ------(((((.((((.(((.....)))........)))))))))(((((((((((.((((....))...(((((....)))..))....................))))))))))))). ( -28.50) >DroSec_CAF1 63558 114 + 1 ------GGGGCAGGGGGAAUCCCUGGCUAAAUUUAAUUUCGCCCCAAAGAAAAAUCACACGAAAUCGACACCUGCCGUGGCGCCGGAAAAUAAUCAAAAUAAAAUAUGGAUUUUUCUUUG ------(((((.((((....))))((............)))))))(((((((((((.((((....))...(((((....)))..))....................))))))))))))). ( -29.30) >DroSim_CAF1 59839 114 + 1 ------GGGGCAGGGGGAAUCCCUGGCUAAAUUUAAUUUCGCCCCAAAGAAAAAUCACACGAAAUCGACACCUGCCGUGGCGCCGGAAAAUAAUCAAAAUAAAAUAUGGAUUUUUCUUUG ------(((((.((((....))))((............)))))))(((((((((((.((((....))...(((((....)))..))....................))))))))))))). ( -29.30) >DroEre_CAF1 58915 119 + 1 GGGCAGGGGGCAGGGAAAAUCCCAGGCUAAAUUUAAUUUCGCCCCAAAGAAAAAUCACACGAAAUCGACACCUGCCGUGGCGCCGGAAAAUAAUCAAAAUAAAAUAUGGAUU-UUCUUUG .((((((((((.(((.....))).((............)))))))..............((....))....)))))........(((((((..(((..........))))))-))))... ( -26.20) >DroYak_CAF1 60602 114 + 1 ------GGGACAGGGGAAAUCCCAGGCUAAAUUUAAUUUCGCCCCAAAGAAAAAUCACACGAAAUCGACACCUGCCGUGGCACCGGAAAAUAAUCAAAAUAAAAUAUGGAUUUUUCUUUG ------((.....(((....))).(((.((((...)))).)))))(((((((((((.((((....))...(((((....)))..))....................))))))))))))). ( -26.20) >consensus ______GGGGCAGGGGGAAUCCCUGGCUAAAUUUAAUUUCGCCCCAAAGAAAAAUCACACGAAAUCGACACCUGCCGUGGCGCCGGAAAAUAAUCAAAAUAAAAUAUGGAUUUUUCUUUG ......(((((.(((.....))).((............)))))))(((((((((((.((((....))...(((((....)))..))....................))))))))))))). (-25.66 = -26.10 + 0.44)



| Location | 11,324,326 – 11,324,440 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.85 |
| Mean single sequence MFE | -34.72 |
| Consensus MFE | -32.86 |
| Energy contribution | -32.58 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.95 |
| SVM decision value | 5.00 |
| SVM RNA-class probability | 0.999967 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11324326 114 - 27905053 CAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGCGAAAUUAAAUUUAGCCAGGGACUCCUCCUGCCCC------ (((((((((((((((..........((......))(((((((......))))))).....))).))))))))))))(((((((......))).....((((....)))))))).------ ( -34.10) >DroSec_CAF1 63558 114 - 1 CAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGCGAAAUUAAAUUUAGCCAGGGAUUCCCCCUGCCCC------ (((((((((((((((..........((......))(((((((......))))))).....))).))))))))))))(((((((......)))....((((.....)))))))).------ ( -36.30) >DroSim_CAF1 59839 114 - 1 CAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGCGAAAUUAAAUUUAGCCAGGGAUUCCCCCUGCCCC------ (((((((((((((((..........((......))(((((((......))))))).....))).))))))))))))(((((((......)))....((((.....)))))))).------ ( -36.30) >DroEre_CAF1 58915 119 - 1 CAAAGAA-AAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGCGAAAUUAAAUUUAGCCUGGGAUUUUCCCUGCCCCCUGCCC (((((((-((..(((((........((......))(((((((......))))))).....)))))..)))))))))(((((((......))).((..(((.....))).)).....)))) ( -33.80) >DroYak_CAF1 60602 114 - 1 CAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGUGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGCGAAAUUAAAUUUAGCCUGGGAUUUCCCCUGUCCC------ (((((((((((((((..........((......))(((..((......))..))).....))).))))))))))))((((.((((...)))).))))(((((.......)))))------ ( -33.10) >consensus CAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGCGAAAUUAAAUUUAGCCAGGGAUUCCCCCUGCCCC______ (((((((((((((((..........((......))(((((((......))))))).....))).))))))))))))(((((((......))).....(((.....))).))))....... (-32.86 = -32.58 + -0.28)



| Location | 11,324,360 – 11,324,479 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.99 |
| Mean single sequence MFE | -36.44 |
| Consensus MFE | -32.44 |
| Energy contribution | -32.68 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979159 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11324360 119 - 27905053 UCACCGUCUCUAACUC-GAUUCGCCUUCGGGUGCGUGGAACAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGC ...((.........((-.((.((((....)))).)).)).(((((((((((((((..........((......))(((((((......))))))).....))).)))))))))))))).. ( -38.90) >DroSec_CAF1 63592 119 - 1 UCACCGUCUCUAACUC-GAUUCGCCUUCGGGUGCGUGGAACAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGC ...((.........((-.((.((((....)))).)).)).(((((((((((((((..........((......))(((((((......))))))).....))).)))))))))))))).. ( -38.90) >DroSim_CAF1 59873 119 - 1 UCACCGUCUCUAACUC-GAUUCGCCUUCGGGUGCGUGGAACAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGC ...((.........((-.((.((((....)))).)).)).(((((((((((((((..........((......))(((((((......))))))).....))).)))))))))))))).. ( -38.90) >DroEre_CAF1 58955 119 - 1 UCACCGUCUCUAACUGGGAUUCGCCUUCGGGUGCGUGGAACAAAGAA-AAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGC ((((.(((((.....))))).((((....)))).))))..(((((((-((..(((((........((......))(((((((......))))))).....)))))..))))))))).... ( -36.60) >DroYak_CAF1 60636 119 - 1 UCAGCGUCUCUAACUC-GAUUCGCCUUCGGCUGCGUGGAACAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGUGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGC ...(((..((......-))..))).(((.((...)).)))(((((((((((((((..........((......))(((..((......))..))).....))).)))))))))))).... ( -28.90) >consensus UCACCGUCUCUAACUC_GAUUCGCCUUCGGGUGCGUGGAACAAAGAAAAAUCCAUAUUUUAUUUUGAUUAUUUUCCGGCGCCACGGCAGGUGUCGAUUUCGUGUGAUUUUUCUUUGGGGC .....((((.........((.((((....)))).))....(((((((((((((((..........((......))(((((((......))))))).....))).)))))))))))))))) (-32.44 = -32.68 + 0.24)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:19:44 2006