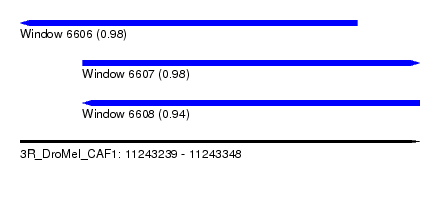

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,243,239 – 11,243,348 |
| Length | 109 |
| Max. P | 0.984863 |

| Location | 11,243,239 – 11,243,331 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 75.89 |
| Mean single sequence MFE | -25.21 |
| Consensus MFE | -21.46 |
| Energy contribution | -21.07 |
| Covariance contribution | -0.39 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978769 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11243239 92 - 27905053 CUCCUCAAAG-GGUUGUGAAAUGUCGACUA-UU-AUCUACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUCG--CG-GCA-GCGGA--UCA--A-AAA (((.((((((-((((((((.(((..((...-..-.))....))).)))))))).)))))).)))........((.(((--(.-...-)))).--)).--.-... ( -27.30) >DroVir_CAF1 2376 94 - 1 CUCUUCAAGG-GGUUGUGAAAUGUUGC-CG-UU-AUCAAGUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUUG--CUUAAAUUCAUUGCCCU--C-GU- (((.((((((-((((((((.(((....-..-..-.......))).)))))))).)))))).)))........(((((.--....)))))........--.-..- ( -23.16) >DroEre_CAF1 2449 93 - 1 CUCCUCAAAG-GGUUGUGAAAUGUCGACUA-UU-AUCUACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUUU--AA-AAA-GCGGG--UUA--AGAAA (((.((((((-((((((((.(((..((...-..-.))....))).)))))))).)))))).)))...(.(((......--..-...-))).)--...--..... ( -22.80) >DroWil_CAF1 2571 99 - 1 CUCUUCAAGGGGGUUGUGAAAUGUUGCCAUUUUAUUCAACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAGUAUUCCU-GCA-ACAGUAUUCG--A-AAU (((.((((((.((((((((...((((..........)))).....)))))))).)))))).))).......((((((((...-...-..))))))))--.-... ( -26.40) >DroMoj_CAF1 2368 93 - 1 CUCUUCAAGG-GGUUGUGAAAUGUAGC-CA-UU-AUCAAGUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGUGAAUUG--CUUUCA-GCAUUGUCCC--A-UC- (((.((((((-((((((((.(((....-..-..-.......))).)))))))).)))))).)))....(((.((..((--(.....-)))...)).)--)-).- ( -25.06) >DroAna_CAF1 2480 92 - 1 CUCCUCAAAG-GGCUGUGAAAUGUUG-CCA-UU-AUCUACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGGGAAUCG-----UCA-GAAAA--CCGUCAGAAA (((.((((((-((((((((.(((...-...-..-.......))).)))))))).)))))).)))((((.(((...((.-----...-))...--)))...)))) ( -26.56) >consensus CUCCUCAAAG_GGUUGUGAAAUGUUGCCCA_UU_AUCAACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUUG__CU_ACA_GCAGU__CCA__A_AAA (((.((((((.((((((((.(((..................))).)))))))).)))))).)))........................................ (-21.46 = -21.07 + -0.39)

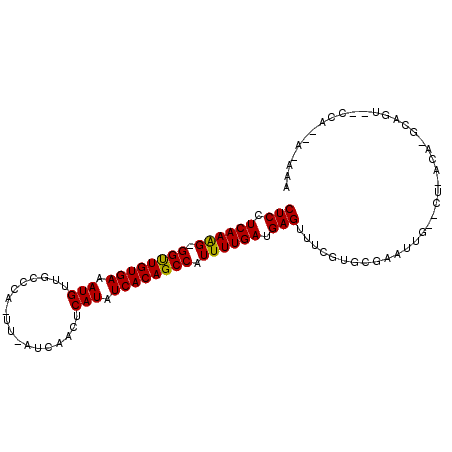

| Location | 11,243,256 – 11,243,348 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.10 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -21.22 |
| Energy contribution | -21.89 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.99 |
| SVM RNA-class probability | 0.984863 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11243256 92 + 27905053 --CGAUUCGCACGAAACUCAUCAAAAUGGCUGUGAUAUGAGUAGAU-AA-UAGUCGACAUUUCACAACC-CUUUGAGGAGUUACGUAAGAAUU--AUGG--------------------- --.(((((..(((.(((((.(((((..((.(((((.(((....(((-..-..)))..))).))))).))-.))))).))))).)))..)))))--....--------------------- ( -28.00) >DroVir_CAF1 2396 94 + 1 --CAAUUCGCACGAAACUCAUCAAAAUGGCUGUGAUAUGACUUGAU-AA-CG-GCAACAUUUCACAACC-CCUUGAAGAGUUACGCAAGAAUUGAGUGGG-------------------- --((((((...((.(((((.((((...((.(((((.(((..(((..-..-..-.)))))).))))).))-..)))).))))).))...))))))......-------------------- ( -25.60) >DroGri_CAF1 2539 94 + 1 --CAAUUCGCACGAAACUCAUCAAAAUGGCUGUGAUAUGACUUGAU-AA-UA-CCCACAUUUCACAACC-CCUUGAAGAGUUACGCAAGAAUUGGGUGGG-------------------- --((((((...((.(((((.((((...((.(((((.(((...((..-..-..-..))))).))))).))-..)))).))))).))...))))))......-------------------- ( -25.00) >DroWil_CAF1 2590 108 + 1 GAAUACUCGCACGAAACUCAUCAAAAUGGCUGUGAUAUGAGUUGAAUAAAAUGGCAACAUUUCACAACCCCCUUGAAGAGUUACGUAAGAAAUU-CCAU--------CUUUUGCCGG--- ((((..((..(((.(((((.((((...((.((((((((.......))).((((....))))))))).))...)))).))))).)))..)).)))-)...--------..........--- ( -24.10) >DroMoj_CAF1 2387 94 + 1 --CAAUUCACACGAAACUCAUCAAAAUGGCUGUGAUAUGACUUGAU-AA-UG-GCUACAUUUCACAACC-CCUUGAAGAGUUACGCAAGAAUUGAAUGGG-------------------- --((((((...((.(((((.((((...((.(((((.(((.((....-..-.)-)...))).))))).))-..)))).))))).))...))))))......-------------------- ( -23.70) >DroAna_CAF1 2498 114 + 1 --CGAUUCCCACGAAACUCAUCAAAAUGGCUGUGAUAUGAGUAGAU-AA-UGG-CAACAUUUCACAGCC-CUUUGAGGAGUUACGUAAGAACUCGAUGGAUCGUCCUUUUUUGAGGGGAU --....(((((((.(((((.(((((..((((((((...........-((-((.-...))))))))))))-.))))).))))).))).....(((((.((.....))....))))))))). ( -41.40) >consensus __CAAUUCGCACGAAACUCAUCAAAAUGGCUGUGAUAUGACUUGAU_AA_UG_GCAACAUUUCACAACC_CCUUGAAGAGUUACGCAAGAAUUG_AUGGG____________________ ..((((((...((.(((((.((((...((.(((((.(((..................))).))))).))...)))).))))).))...)))))).......................... (-21.22 = -21.89 + 0.67)


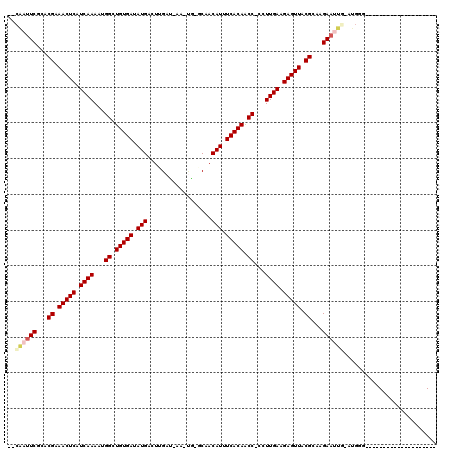
| Location | 11,243,256 – 11,243,348 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.10 |
| Mean single sequence MFE | -33.45 |
| Consensus MFE | -31.15 |
| Energy contribution | -30.45 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.17 |
| Mean z-score | -4.25 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.941281 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11243256 92 - 27905053 ---------------------CCAU--AAUUCUUACGUAACUCCUCAAAG-GGUUGUGAAAUGUCGACUA-UU-AUCUACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUCG-- ---------------------....--.((((.((((.(((((.((((((-((((((((.(((..((...-..-.))....))).)))))))).)))))).))))).)))).))))..-- ( -30.90) >DroVir_CAF1 2396 94 - 1 --------------------CCCACUCAAUUCUUGCGUAACUCUUCAAGG-GGUUGUGAAAUGUUGC-CG-UU-AUCAAGUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUUG-- --------------------......((((((.((((.(((((.((((((-((((((((.(((....-..-..-.......))).)))))))).)))))).))))).)))).))))))-- ( -32.46) >DroGri_CAF1 2539 94 - 1 --------------------CCCACCCAAUUCUUGCGUAACUCUUCAAGG-GGUUGUGAAAUGUGGG-UA-UU-AUCAAGUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUUG-- --------------------......((((((.((((.(((((.((((((-((((((((..(((((.-(.-..-....).))))))))))))).)))))).))))).)))).))))))-- ( -33.80) >DroWil_CAF1 2590 108 - 1 ---CCGGCAAAAG--------AUGG-AAUUUCUUACGUAACUCUUCAAGGGGGUUGUGAAAUGUUGCCAUUUUAUUCAACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAGUAUUC ---..........--------...(-((((((.((((.(((((.((((((.((((((((...((((..........)))).....)))))))).)))))).))))).)))).))).)))) ( -31.30) >DroMoj_CAF1 2387 94 - 1 --------------------CCCAUUCAAUUCUUGCGUAACUCUUCAAGG-GGUUGUGAAAUGUAGC-CA-UU-AUCAAGUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGUGAAUUG-- --------------------......((((((.((((.(((((.((((((-((((((((.(((....-..-..-.......))).)))))))).)))))).))))).)))).))))))-- ( -32.46) >DroAna_CAF1 2498 114 - 1 AUCCCCUCAAAAAAGGACGAUCCAUCGAGUUCUUACGUAACUCCUCAAAG-GGCUGUGAAAUGUUG-CCA-UU-AUCUACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGGGAAUCG-- ....(((......))).(((....))).(((((((((.(((((.((((((-((((((((.(((...-...-..-.......))).)))))))).)))))).))))).)))))))))..-- ( -39.76) >consensus ____________________CCCAU_CAAUUCUUACGUAACUCUUCAAGG_GGUUGUGAAAUGUUGC_CA_UU_AUCAACUCAUAUCACAGCCAUUUUGAUGAGUUUCGUGCGAAUUG__ ............................((((.((((.(((((.((((((.((((((((.(((..................))).)))))))).)))))).))))).)))).)))).... (-31.15 = -30.45 + -0.69)
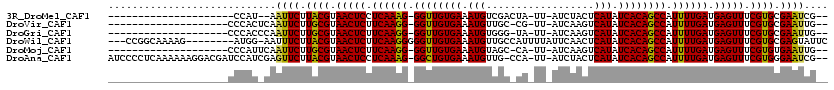

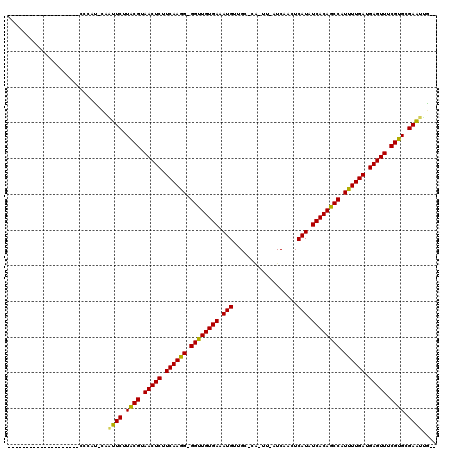
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:18:48 2006