
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,196,153 – 11,196,256 |
| Length | 103 |
| Max. P | 0.992039 |

| Location | 11,196,153 – 11,196,256 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 67.91 |
| Mean single sequence MFE | -32.86 |
| Consensus MFE | -19.92 |
| Energy contribution | -20.65 |
| Covariance contribution | 0.73 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.61 |
| SVM decision value | 2.30 |
| SVM RNA-class probability | 0.992039 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11196153 103 + 27905053 AUCUCCACCACGUCUUGGGUUUG---CUCU---UCC-----UCUUCAGGCUCCAUGUG----AGACUUGCAUUGCGUCCAACAUGCCUCUCUGAUGUUGGUCGCCUUGCAAGUCCUAC-- ........(((((...(((((((---....---...-----....))))))).)))))----.((((((((..(((.(((((((.........))))))).)))..))))))))....-- ( -36.00) >DroVir_CAF1 6881 102 + 1 AGCACUGCCAGCUGGUCAAC---GGGCAAAGUAUUC-----U------CCUGCAACGG----GGCCAUGCAAUGGGUCCAGCAUGUAUCGGCUAUGUUGGGCCCCCUGCAAGACCACUGC ........(((.(((((...---.(((..((....)-----)------((((...)))----)))).((((..(((((((((((((....)).)))))))))))..)))).)))))))). ( -40.90) >DroSec_CAF1 6765 103 + 1 AUCUCCACCACGUCUUGGGCUUG---CUCU---UUC-----UCUUCCGGCUCCAUGUG----AGACUUGCAUUGCGUCCAACAGGCCUCUCUGAUGUCGGUCGCCUUGCAAGUCCUAC-- ........(((((...(((((..---....---...-----......))))).)))))----.((((((((..(((.((.(((...........))).)).)))..))))))))....-- ( -30.06) >DroSim_CAF1 5899 103 + 1 AUCUCUACUACGUCUUGGGUUUG---CUCU---UUC-----UCUUCUGGCUCCAUGUG----AGACUUGCAUUGCGUCCAACAGGCCUCUCUGAUGUCGGUCGCCUUGCAAGUCCUAC-- .................((...(---((..---...-----......))).))..(((----.((((((((..(((.((.(((...........))).)).)))..)))))))).)))-- ( -26.40) >DroYak_CAF1 6217 103 + 1 ACCUCCACCAAGUCCUGGAUGUG---CUCU---UUC-----AUUGGUGGCACGAUGUG----AGACUUGCAUUGCGUCCAACAUGCCUCUCUGAUGUUGGUCGCCUUGCAAGUCCUAC-- ....(((((((.....(((....---.)))---...-----.)))))))......(((----.((((((((..(((.(((((((.........))))))).)))..)))))))).)))-- ( -39.90) >DroAna_CAF1 6486 112 + 1 AUCUCCACCAAGUGGUUGGGAUU---CUCA---ACCUUCUCCAUCAUGCCUUCAUCUGUCGUGGGCUUGCAGUGCGUCCAACUGAUCUGUCUGAUGUUGGUCACCUCACAAGUUUCAC-- .....(((((((((((((((...---))))---))).......(((((.(.......).))))))))))..))).(.(((((..(((.....)))))))).)................-- ( -23.90) >consensus AUCUCCACCACGUCUUGGGUUUG___CUCU___UUC_____UCUUCUGGCUCCAUGUG____AGACUUGCAUUGCGUCCAACAGGCCUCUCUGAUGUUGGUCGCCUUGCAAGUCCUAC__ ...............................................................((((((((..(((.((((((...........)))))).)))..))))))))...... (-19.92 = -20.65 + 0.73)



| Location | 11,196,153 – 11,196,256 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 67.91 |
| Mean single sequence MFE | -33.37 |
| Consensus MFE | -22.85 |
| Energy contribution | -22.13 |
| Covariance contribution | -0.72 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.68 |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.970937 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11196153 103 - 27905053 --GUAGGACUUGCAAGGCGACCAACAUCAGAGAGGCAUGUUGGACGCAAUGCAAGUCU----CACAUGGAGCCUGAAGA-----GGA---AGAG---CAAACCCAAGACGUGGUGGAGAU --((..((((((((..(((.((((((((......).))))))).)))..)))))))).----.)).....((((.....-----...---)).)---)....(((........))).... ( -34.30) >DroVir_CAF1 6881 102 - 1 GCAGUGGUCUUGCAGGGGGCCCAACAUAGCCGAUACAUGCUGGACCCAUUGCAUGGCC----CCGUUGCAGG------A-----GAAUACUUUGCCC---GUUGACCAGCUGGCAGUGCU ((((((.(((((((((((((((((.....(((........))).....)))...))))----)).)))))))------)-----...))).(((((.---(((....))).)))))))). ( -37.70) >DroSec_CAF1 6765 103 - 1 --GUAGGACUUGCAAGGCGACCGACAUCAGAGAGGCCUGUUGGACGCAAUGCAAGUCU----CACAUGGAGCCGGAAGA-----GAA---AGAG---CAAGCCCAAGACGUGGUGGAGAU --((..((((((((..(((.((((((...........)))))).)))..)))))))).----.)).(((.(((....).-----(..---....---)..)))))............... ( -31.90) >DroSim_CAF1 5899 103 - 1 --GUAGGACUUGCAAGGCGACCGACAUCAGAGAGGCCUGUUGGACGCAAUGCAAGUCU----CACAUGGAGCCAGAAGA-----GAA---AGAG---CAAACCCAAGACGUAGUAGAGAU --((..((((((((..(((.((((((...........)))))).)))..)))))))).----.)).(((.(((......-----...---.).)---)....)))............... ( -28.40) >DroYak_CAF1 6217 103 - 1 --GUAGGACUUGCAAGGCGACCAACAUCAGAGAGGCAUGUUGGACGCAAUGCAAGUCU----CACAUCGUGCCACCAAU-----GAA---AGAG---CACAUCCAGGACUUGGUGGAGGU --((..((((((((..(((.((((((((......).))))))).)))..)))))))).----.))(((...(((((((.-----...---.((.---....))......))))))).))) ( -40.30) >DroAna_CAF1 6486 112 - 1 --GUGAAACUUGUGAGGUGACCAACAUCAGACAGAUCAGUUGGACGCACUGCAAGCCCACGACAGAUGAAGGCAUGAUGGAGAAGGU---UGAG---AAUCCCAACCACUUGGUGGAGAU --......((((((..(((.((((((((.....)))..))))).)))..)))))).((((..((.(((....)))..)).....(((---((..---.....))))).....)))).... ( -27.60) >consensus __GUAGGACUUGCAAGGCGACCAACAUCAGAGAGGCAUGUUGGACGCAAUGCAAGUCU____CACAUGGAGCCAGAAGA_____GAA___AGAG___CAAACCCAAGACGUGGUGGAGAU .....(((((((((..(((.((((((((.....))..)))))).)))..))))))))).............................................................. (-22.85 = -22.13 + -0.72)

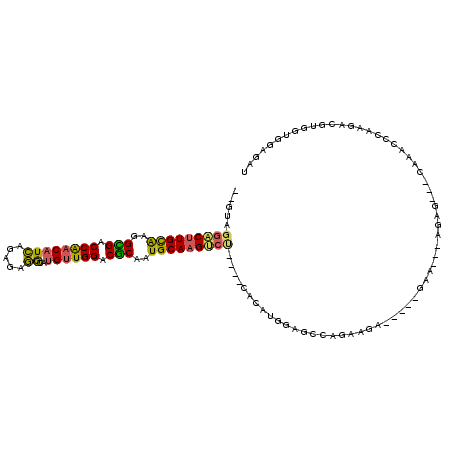

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:18:04 2006