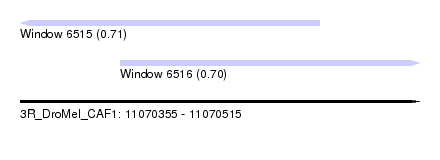
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 11,070,355 – 11,070,515 |
| Length | 160 |
| Max. P | 0.714735 |

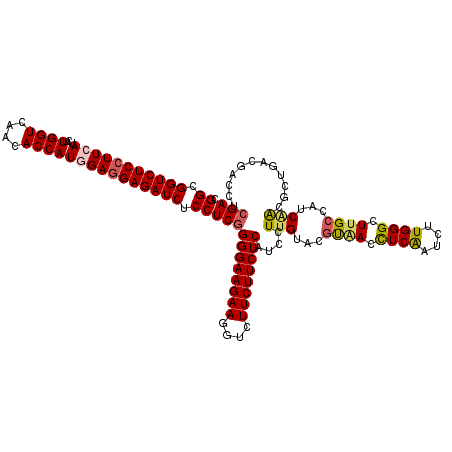
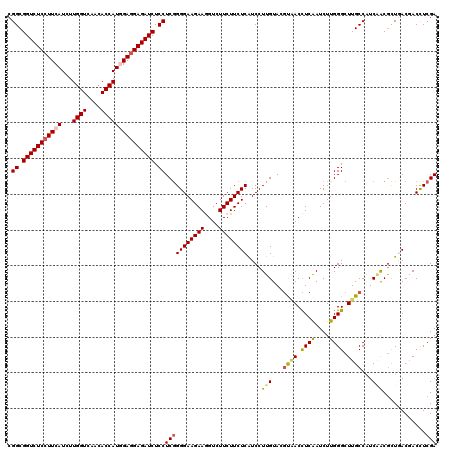
| Location | 11,070,355 – 11,070,475 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.89 |
| Mean single sequence MFE | -41.15 |
| Consensus MFE | -33.40 |
| Energy contribution | -33.60 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.714735 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11070355 120 - 27905053 CGGCGGUCUCCUUCAUCUUGGUAAGCACCAUCGAAGAGAUCUCCUCGGGGAAGAAGGUCUUCUUCUCGUCCUUGUAGGUCACCUCGAUCUUGGGCUUGCCAUCGGCGCUGACCACCUCGA .((.((((..(.((....((((((((.(((((((.(.(((((...(((((((((((.....)))))..)))))).))))).).))))...)))))))))))..)).)..)))).)).... ( -45.60) >DroVir_CAF1 2484 120 - 1 CGGCGGUCUCCUUCAUUUUGGUCAACACCAUGGAGGAGAUCUCCUCGGGGAAGAAAGUCUUCUUCUCAUCCUUGUACGUAACUUCAAUCUUGGGCUUGCCAUCAACGCUAACGACCUCGA .((.(((((((((((...((((....))))))))))))))).))(((((((((((....))))))))....(((...((((.((((....)))).))))...)))......)))...... ( -37.10) >DroGri_CAF1 278 120 - 1 CGGCGGUCUCCUUCAUUUUGGUCAACACCAUGGAGGAGAUCUCCUCGGGGAAGAAGGUCUUCUUCUCGUCCUUGUACGUAACCUCAAUCUUGGGCUUGCCAUCAACGCUGACAACUUCGA .((.(((((((((((...((((....))))))))))))))).))(((((((((((....))))))))(((.(((...((((.((((....)))).))))...)))....))).....))) ( -41.60) >DroWil_CAF1 494 120 - 1 CGGCGGUCUCCUUCAUUUUGGUCAACACCAUGGAGGAGAUCUCCUCGGGGAAGAAGGUCUUCUUCUCAUCCUUGUAGACAACUUCAAUCUUGGGCUUGCCAUCUACACUGACGACCUCGA .((.(((((((((((...((((....))))))))))))))).))((((((..((((.....))))((.((..(((((((((........)))((....)).))))))..)).)))))))) ( -42.50) >DroMoj_CAF1 278 120 - 1 CGGCGGUCUCCUUCAUCUUGGUCAACACCAUGGAGGAGAUCUCCUCAGGGAAGAAAGUCUUCUUCUCAUCCUUGUACGUAACCUCGAUCUUGGGCUUGCCAUCAACGUUAACGACCUCGA .((((((((((((((...((((....)))))))))))))))..((((((((((((....)))))))).........((......))....))))...)))...........((....)). ( -37.30) >DroAna_CAF1 278 120 - 1 CGGCGGUCUCCUUCAUCUUGGUGAGGACCAUAGAGGAGAUCUCCUCGGGGAAGAAGGUCUUCUUCUCAUCCUUGUAGGUGACCUCGAUCUUGGGCUUGCCGUCAACGCUGACAACCUCGA .((.(((((((((.....((((....))))..))))))))).))(((((((((((((((.(((...((....)).))).)))))...)))).((....))((((....))))..)))))) ( -42.80) >consensus CGGCGGUCUCCUUCAUCUUGGUCAACACCAUGGAGGAGAUCUCCUCGGGGAAGAAGGUCUUCUUCUCAUCCUUGUACGUAACCUCAAUCUUGGGCUUGCCAUCAACGCUGACGACCUCGA .((.(((((((((((...((((....))))))))))))))).))(((((((((((....))))))))....(((...((((.((((....)))).))))...)))............))) (-33.40 = -33.60 + 0.20)

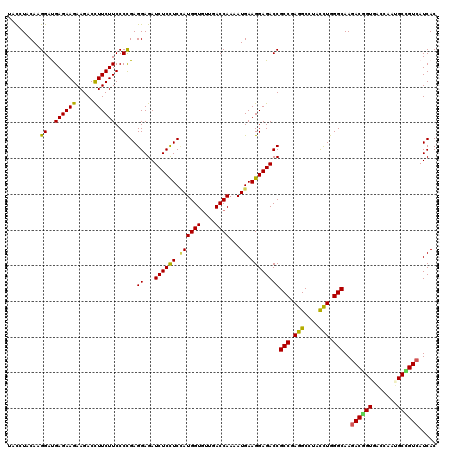
| Location | 11,070,395 – 11,070,515 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.11 |
| Mean single sequence MFE | -38.22 |
| Consensus MFE | -35.85 |
| Energy contribution | -34.97 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.697828 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 11070395 120 + 27905053 GACCUACAAGGACGAGAAGAAGACCUUCUUCCCCGAGGAGAUCUCUUCGAUGGUGCUUACCAAGAUGAAGGAGACCGCCGAGGCCUAUCUGGGCAAGACUGUGACCAACGCGGUCAUCAC ..(((....((..((((((.....))))))..)).))).(.((((((((.((((....))))...)))).)))).)(((.(((....))).)))..(((((((.....)))))))..... ( -37.40) >DroVir_CAF1 2524 120 + 1 UACGUACAAGGAUGAGAAGAAGACUUUCUUCCCCGAGGAGAUCUCCUCCAUGGUGUUGACCAAAAUGAAGGAGACCGCCGAGGCGUACCUGGGCAAGACGGUUACCAAUGCCGUCAUCAC .........((..((((((.....))))))..))..((...((((((.((((((....))))...)).))))))))(((.(((....))).)))..((((((.......))))))..... ( -39.40) >DroGri_CAF1 318 120 + 1 UACGUACAAGGACGAGAAGAAGACCUUCUUCCCCGAGGAGAUCUCCUCCAUGGUGUUGACCAAAAUGAAGGAGACCGCCGAGGCGUAUCUGGGCAAGACGGUGACCAAUGCCGUGAUCAC .........((..((((((.....))))))..))(((((....)))))(((((((((((((........(....).(((.(((....))).))).....)))...))))))))))..... ( -39.00) >DroWil_CAF1 534 120 + 1 UGUCUACAAGGAUGAGAAGAAGACCUUCUUCCCCGAGGAGAUCUCCUCCAUGGUGUUGACCAAAAUGAAGGAGACCGCCGAGGCCUACUUGGGCAAGACAGUGGCCAAUGCUGUCAUCAC ((((((..(((....((((((....)))))).((..((.(.((((((.((((((....))))...)).)))))).).))..)))))...)))))).(((((((.....)))))))..... ( -40.20) >DroMoj_CAF1 318 120 + 1 UACGUACAAGGAUGAGAAGAAGACUUUCUUCCCUGAGGAGAUCUCCUCCAUGGUGUUGACCAAGAUGAAGGAGACCGCCGAGGCGUACCUGGGCAAGACGGUUACCAAUGCCGUCAUCAC ......((.(((.((((.......)))).))).)).((...((((((.((((((....))))...)).))))))))(((.(((....))).)))..((((((.......))))))..... ( -39.60) >DroAna_CAF1 318 120 + 1 CACCUACAAGGAUGAGAAGAAGACCUUCUUCCCCGAGGAGAUCUCCUCUAUGGUCCUCACCAAGAUGAAGGAGACCGCCGAGGCCUAUCUGGGCAAGACUGUGACCAAUGCCGUCAUCAC ..(((....((..((((((.....))))))..)).))).(.(((((((..((((....))))....).)))))).)(((.(((....))).)))..(((.((.......)).)))..... ( -33.70) >consensus UACCUACAAGGAUGAGAAGAAGACCUUCUUCCCCGAGGAGAUCUCCUCCAUGGUGUUGACCAAAAUGAAGGAGACCGCCGAGGCCUACCUGGGCAAGACGGUGACCAAUGCCGUCAUCAC .........((..((((((.....))))))..))..((...((((((.((((((....))))...)).))))))))(((.(((....))).)))..((((((.......))))))..... (-35.85 = -34.97 + -0.88)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:17:18 2006