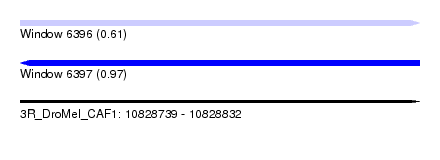

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 10,828,739 – 10,828,832 |
| Length | 93 |
| Max. P | 0.969650 |

| Location | 10,828,739 – 10,828,832 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 91.40 |
| Mean single sequence MFE | -24.04 |
| Consensus MFE | -18.30 |
| Energy contribution | -18.90 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.613336 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10828739 93 + 27905053 AAUUCGAGUGUGCGCUCUAGCGUAAAGGCCAAACUGCACCUGCACCUCAAAAAGAUGUAGUAAUCUGUGCACAUCAGUUACAAACUAUCGGCC-- ..........(((((....)))))..((((....((((((((((.((.....)).)))))......)))))....(((.....)))...))))-- ( -21.80) >DroSec_CAF1 98216 93 + 1 GAUUCGAGUGUGCGCUCUAGCGUAAAGGCCAAACUGCACCUGCACCUCAAAAAGAUGCAGUAAUCUGUGCACAUUAGUUACAAACUAUCGGCC-- ..........(((((....)))))..((((....((((((((((.((.....)).)))))......)))))...((((.....))))..))))-- ( -24.90) >DroSim_CAF1 104330 93 + 1 GAUUCGAGUGUGCGCUCUAGCGUAAAGGCCAAACUGCACCUGCACCUCAAAAAGAUGCAGUAAUCUGUGCACAUCAGUAACAAACUAUCGGCC-- ..........(((((....)))))..((((....((((((((((.((.....)).)))))......)))))....(((.....)))...))))-- ( -24.00) >DroEre_CAF1 100661 93 + 1 AAUUCGAGUGUGCGCUCUAGCGUAAAGGCCAAAAUGCACCUGCACCUCAAAAAUAUGCAGUAAUCUGUGCACAUCAGUUACAAACUC--GGAUGU .((((((((.(((((....)))))..........((((((((((...........)))))......)))))............))))--)))).. ( -24.20) >DroYak_CAF1 103762 93 + 1 AAUUCGAGUGUGCGCUCUAGCGUAAAGGCCAAACUGCACCUGCACCUCAAACAGAUGCAGUAAUCUUUGCACAUCAGUUACAAACUG--GAAUGU .((((..((((((((....))..(((((....((((((.(((.........))).))))))..)))))))))))((((.....))))--)))).. ( -25.30) >consensus AAUUCGAGUGUGCGCUCUAGCGUAAAGGCCAAACUGCACCUGCACCUCAAAAAGAUGCAGUAAUCUGUGCACAUCAGUUACAAACUAUCGGCC__ ..........(((((....)))))..((((....((((((((((.((.....)).)))))......)))))....(((.....)))...)))).. (-18.30 = -18.90 + 0.60)


| Location | 10,828,739 – 10,828,832 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 91.40 |
| Mean single sequence MFE | -29.18 |
| Consensus MFE | -24.40 |
| Energy contribution | -25.00 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969650 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10828739 93 - 27905053 --GGCCGAUAGUUUGUAACUGAUGUGCACAGAUUACUACAUCUUUUUGAGGUGCAGGUGCAGUUUGGCCUUUACGCUAGAGCGCACACUCGAAUU --...(((((((((((.((....))..))))))))((.((((((...)))))).))((((..((((((......))))))..))))..))).... ( -24.30) >DroSec_CAF1 98216 93 - 1 --GGCCGAUAGUUUGUAACUAAUGUGCACAGAUUACUGCAUCUUUUUGAGGUGCAGGUGCAGUUUGGCCUUUACGCUAGAGCGCACACUCGAAUC --...(((((((((((.((....))..))))))))(((((((((...)))))))))((((..((((((......))))))..))))..))).... ( -30.90) >DroSim_CAF1 104330 93 - 1 --GGCCGAUAGUUUGUUACUGAUGUGCACAGAUUACUGCAUCUUUUUGAGGUGCAGGUGCAGUUUGGCCUUUACGCUAGAGCGCACACUCGAAUC --...((((((((((((((....))).))))))))(((((((((...)))))))))((((..((((((......))))))..))))..))).... ( -31.50) >DroEre_CAF1 100661 93 - 1 ACAUCC--GAGUUUGUAACUGAUGUGCACAGAUUACUGCAUAUUUUUGAGGUGCAGGUGCAUUUUGGCCUUUACGCUAGAGCGCACACUCGAAUU .....(--(((((((((.((((((((((........))))))))....)).)))))((((.(((((((......))))))).))))))))).... ( -28.70) >DroYak_CAF1 103762 93 - 1 ACAUUC--CAGUUUGUAACUGAUGUGCAAAGAUUACUGCAUCUGUUUGAGGUGCAGGUGCAGUUUGGCCUUUACGCUAGAGCGCACACUCGAAUU ..((((--((((.....)))).(((((........((((((((((.......))))))))))((((((......))))))..)))))...)))). ( -30.50) >consensus __GGCCGAUAGUUUGUAACUGAUGUGCACAGAUUACUGCAUCUUUUUGAGGUGCAGGUGCAGUUUGGCCUUUACGCUAGAGCGCACACUCGAAUU .........(((((((.((....))..))))))).((((((((.....))))))))((((..((((((......))))))..))))......... (-24.40 = -25.00 + 0.60)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:15:19 2006