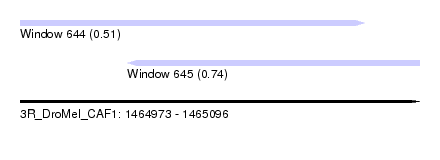
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 1,464,973 – 1,465,096 |
| Length | 123 |
| Max. P | 0.735813 |

| Location | 1,464,973 – 1,465,079 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.51 |
| Mean single sequence MFE | -26.03 |
| Consensus MFE | -22.08 |
| Energy contribution | -22.50 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.85 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.506624 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
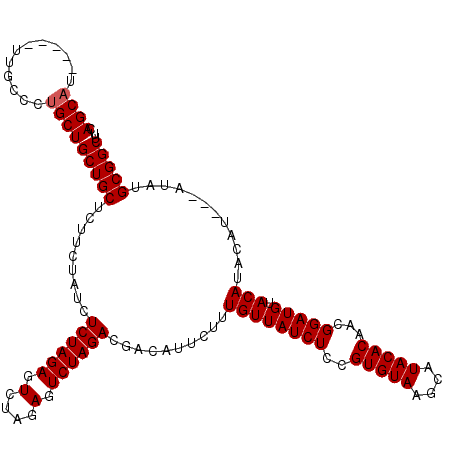
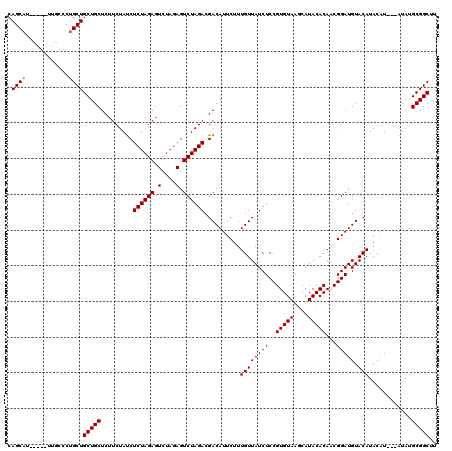
>3R_DroMel_CAF1 1464973 106 + 27905053 CAGC-------UUGCCCUGCUGCUGCUCUUCUAUCUCUAGA-------GUCUAGACGACAUUCUUUGUUAUCUCCGUGUAAUCAUACACAACGGAUGUACAUACUUAUUAUAUGCGGCUU .(((-------.......)))(((((...((((....))))-------(((((((..(((.....)))..)))..(((((....)))))...)))).................))))).. ( -22.40) >DroSec_CAF1 37492 117 + 1 CAGCAUUCAGCUUGCCCUGCUGCUGCUCUUCUAUCUCUAGAGUCUAGAGUCUAGACGACAUUCUUUGUUAUCUCCGUGUAAGCAUACACAGCGGAUGUACAUACAU---AUAUGCGGCUU .((((..((((.......)))).))))..((((.((((((...))))))..)))).((((.....))))....(((((((....))))).(((.((((....))))---...)))))... ( -28.40) >DroSim_CAF1 35777 108 + 1 CAGCAU---------CCUGCUGCUGCUCUUCUACCUCUAGAGUCUAGAGUCUAGACGACAUUCUUUGUUAUCUCCGUGUAAGCAUACACAUCGGAUGUACAUACAU---AUAUGCGGCUU .(((..---------...)))(((((((.((((.((((((...))))))..)))).)).......((((((((..(((((....)))))...))))).))).....---....))))).. ( -27.30) >consensus CAGCAU_____UUGCCCUGCUGCUGCUCUUCUAUCUCUAGAGUCUAGAGUCUAGACGACAUUCUUUGUUAUCUCCGUGUAAGCAUACACAACGGAUGUACAUACAU___AUAUGCGGCUU .((((............))))(((((.........((((((.(....).))))))..........((((((((..(((((....)))))...))))).)))............))))).. (-22.08 = -22.50 + 0.42)

| Location | 1,465,006 – 1,465,096 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 89.93 |
| Mean single sequence MFE | -22.83 |
| Consensus MFE | -19.27 |
| Energy contribution | -19.27 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.735813 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
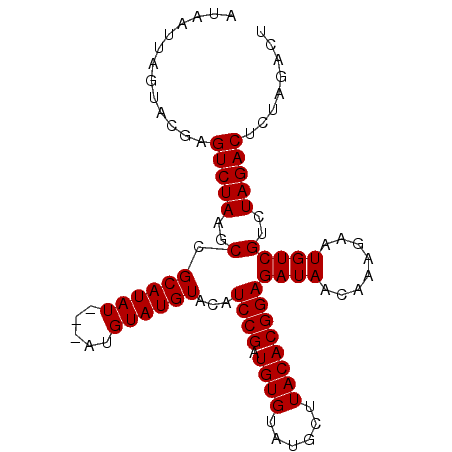
>3R_DroMel_CAF1 1465006 90 - 27905053 AUAAUUAGUACGAGUCUAAGCCGCAUAUAAUAAGUAUGUACAUCCGUUGUGUAUGAUUACACGGAGAUAACAAAGAAUGUCGUCUAGAC-------U ............((((((..(.((((((.....))))))...(((((.(((......))))))))((((........)))))..)))))-------) ( -20.60) >DroSec_CAF1 37532 94 - 1 AUAAUUAGUACAAGUCUAAGCCGCAUAU---AUGUAUGUACAUCCGCUGUGUAUGCUUACACGGAGAUAACAAAGAAUGUCGUCUAGACUCUAGACU ......(((...((((((..(.((((..---.(((.(((...((((.((((......)))))))).))))))....)))).)..))))))....))) ( -22.20) >DroSim_CAF1 35808 94 - 1 AUAAUUAGUACGAGUCUAAGCCGCAUAU---AUGUAUGUACAUCCGAUGUGUAUGCUUACACGGAGAUAACAAAGAAUGUCGUCUAGACUCUAGACU ......(((..(((((((..(.((((..---.(((.(((...((((.((((......)))))))).))))))....)))).)..)))))))...))) ( -25.70) >consensus AUAAUUAGUACGAGUCUAAGCCGCAUAU___AUGUAUGUACAUCCGAUGUGUAUGCUUACACGGAGAUAACAAAGAAUGUCGUCUAGACUCUAGACU .............(((((..(.((((((.....))))))...((((.((((......))))))))((((........)))))..)))))........ (-19.27 = -19.27 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:42:05 2006