
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 10,540,906 – 10,541,040 |
| Length | 134 |
| Max. P | 0.999953 |

| Location | 10,540,906 – 10,541,002 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 93.08 |
| Mean single sequence MFE | -24.78 |
| Consensus MFE | -21.86 |
| Energy contribution | -21.90 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.88 |
| SVM decision value | 2.52 |
| SVM RNA-class probability | 0.994918 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
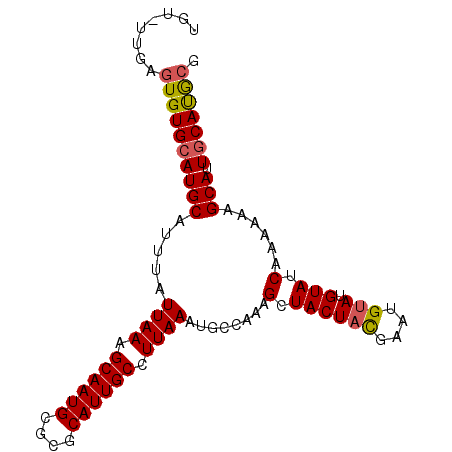
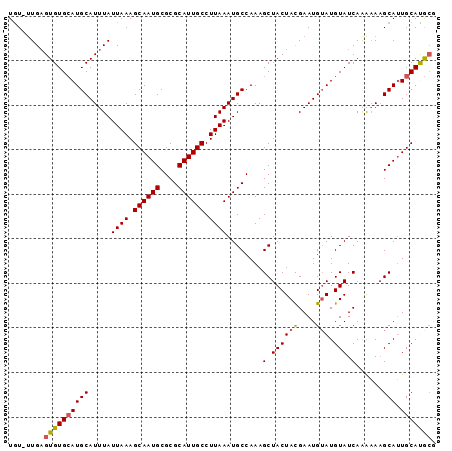
>3R_DroMel_CAF1 10540906 96 + 27905053 UGUGUUGAGUGUGCAUGCAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAAAAAGCAUUGCACGCG ........((((((((((.....((((.((((((....)))))).))))........(.((((((....))).))).)......))).))))))). ( -26.30) >DroSec_CAF1 15853 95 + 1 UGU-UUGAGUGUGCAUGCAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGAAUGUAUCAAAAAAGCAUUGCACGGG .((-(((((((.((..(((((((.....((((((....))))))..)))))))....)))))).))))).(((((.........)))))....... ( -21.70) >DroEre_CAF1 15966 95 + 1 UGU-UUGAGUGUGCAUGCAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAAAAAGCAUUGCAUGCG .((-(((((((.((..(((((((.....((((((....))))))..)))))))....)))))).)))))(((((((............))))))). ( -26.70) >DroYak_CAF1 15980 95 + 1 UGU-UUGAGUGUGCAUGCAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUAUGAAUGUAUGUAUCAGAAAAGCAUUGCAUGCG ..(-((((...((((((((((((((((.((((((....)))))).))))..((....)).....)))))))))))))))))...((((...)))). ( -26.40) >DroAna_CAF1 15742 95 + 1 UAU-CUGUGUGUGAAUGCAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAGUAUGCAAUGCAUACA ...-..((((((..((((((((.((((.((((((....)))))).))))..((....))......))))))))((((.....))))...)))))). ( -22.80) >consensus UGU_UUGAGUGUGCAUGCAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAAAAAGCAUUGCAUGCG ........((((((((((.....((((.((((((....)))))).))))........(.((((((....))).))).)......))).))))))). (-21.86 = -21.90 + 0.04)

| Location | 10,540,906 – 10,541,002 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 93.08 |
| Mean single sequence MFE | -28.48 |
| Consensus MFE | -25.50 |
| Energy contribution | -26.46 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.89 |
| Structure conservation index | 0.90 |
| SVM decision value | 4.82 |
| SVM RNA-class probability | 0.999953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
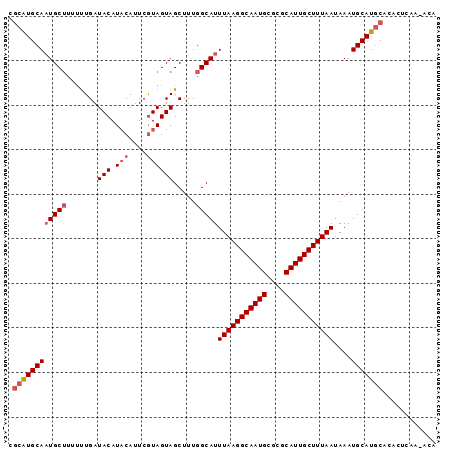
>3R_DroMel_CAF1 10540906 96 - 27905053 CGCGUGCAAUGCUUUUUUGAUACAUACAUUCGUAGUAGCUUUGGCAUUUAAGGCAAUGCGCGCAUUGCUUUAAUAAAUGCAUGCACACUCAACACA .((((((((((((.......(((.(((....)))))).....)))))(((((((((((....)))))))))))....)))))))............ ( -30.80) >DroSec_CAF1 15853 95 - 1 CCCGUGCAAUGCUUUUUUGAUACAUUCAUUCGUAGUAGCUUUGGCAUUUAAGGCAAUGCGCGCAUUGCUUUAAUAAAUGCAUGCACACUCAA-ACA ...(((((.((((....(((.....))).....))))......(((((((((((((((....))))))))...))))))).)))))......-... ( -27.40) >DroEre_CAF1 15966 95 - 1 CGCAUGCAAUGCUUUUUUGAUACAUACAUUCGUAGUAGCUUUGGCAUUUAAGGCAAUGCGCGCAUUGCUUUAAUAAAUGCAUGCACACUCAA-ACA .((((((((((((.......(((.(((....)))))).....)))))(((((((((((....)))))))))))....)))))))........-... ( -31.20) >DroYak_CAF1 15980 95 - 1 CGCAUGCAAUGCUUUUCUGAUACAUACAUUCAUAGUAGCUUUGGCAUUUAAGGCAAUGCGCGCAUUGCUUUAAUAAAUGCAUGCACACUCAA-ACA .((((((((((((.......(((...........))).....)))))(((((((((((....)))))))))))....)))))))........-... ( -28.40) >DroAna_CAF1 15742 95 - 1 UGUAUGCAUUGCAUACUUGAUACAUACAUUCGUAGUAGCUUUGGCAUUUAAGGCAAUGCGCGCAUUGCUUUAAUAAAUGCAUUCACACACAG-AUA ((.(((((((((.(((((((.........))).))))))........(((((((((((....)))))))))))..))))))).)).......-... ( -24.60) >consensus CGCAUGCAAUGCUUUUUUGAUACAUACAUUCGUAGUAGCUUUGGCAUUUAAGGCAAUGCGCGCAUUGCUUUAAUAAAUGCAUGCACACUCAA_ACA .((((((((((((.......(((.(((....)))))).....)))))(((((((((((....)))))))))))....)))))))............ (-25.50 = -26.46 + 0.96)


| Location | 10,540,923 – 10,541,040 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 82.34 |
| Mean single sequence MFE | -23.90 |
| Consensus MFE | -17.78 |
| Energy contribution | -18.22 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.853029 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10540923 117 + 27905053 CAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAAAAAGCAUUGCACGCGUACACUGGAAACAA-AAUGUAAAGGACUCAAGGCAUU-GU .......(((.((((((....)))))).)))((((((..((((..((((..((((((..((((.......))).)..)))))).((....)).-..))))..)).))...))))))-.. ( -26.70) >DroSec_CAF1 15869 118 + 1 CAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGAAUGUAUCAAAAAAGCAUUGCACGGGUACACUGGAAACAA-AAUGUAAAGGAAUCAAGACAUACGG ...........((((((((.(((((......))))))..............(((.....))).....))))))).(...((((.((....)).-..))))..)................ ( -21.40) >DroEre_CAF1 15982 113 + 1 CAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAAAAAGCAUUGCAUGCGUACACUGGAAACAAGAAAGUC-----CACAAGCCAUU-AA ((((((.....((((((....))))))..)))))).....(((........((((((((((((.......))).))))))))).((....))........-----....)))....-.. ( -23.40) >DroYak_CAF1 15996 106 + 1 CAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUAUGAAUGUAUGUAUCAGAAAAGCAUUGCAUGCGUACACUGGAAACAAUAAUGUA--------AAGCUA----- ((((((.....((((((....))))))..))))))....((((....((.(((((((((............))))))))).)).((....))........--------.)))).----- ( -24.40) >DroAna_CAF1 15758 99 + 1 CAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAGUAUGCAAUGCAUACAUGCACUGAAAAAAA--AGGCAG------------------ ......((((.((((((....)))))).)))).((((...(.((((((....))).))).)..((((((...))))))...............--.)))).------------------ ( -23.60) >consensus CAUUUAUUAAAGCAAUGCGCGCAUUGCCUUAAAUGCCAAAGCUACUACGAAUGUAUGUAUCAAAAAAGCAUUGCAUGCGUACACUGGAAACAA_AAUGUA_______CAAGCCAU____ ...........((((((((.(((((......))))))...(.((((((....))).))).)......)))))))..........((....))........................... (-17.78 = -18.22 + 0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:18 2006