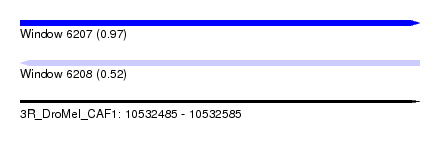

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 10,532,485 – 10,532,585 |
| Length | 100 |
| Max. P | 0.966561 |

| Location | 10,532,485 – 10,532,585 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 86.71 |
| Mean single sequence MFE | -13.74 |
| Consensus MFE | -13.52 |
| Energy contribution | -13.52 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966561 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10532485 100 + 27905053 CUUCGACCAAGCGCUGUAAUAAAUUCGCGCUUUAAA--AAAUAAAGAAAAACAGUAAAAAAUAAAAUGACAAAGA-GGUGGGGGAAAU--AUGAGGAAAAACAAG ((((.(((((((((............))))))....--.............((.............)).......-))).))))....--............... ( -13.82) >DroEre_CAF1 7454 91 + 1 CUUCGACCAAGCGCUGUAAUAAAUUCGCGCUUUAAAAAA---------AAAGAGUAAA-AAUAAAAUGACAAAA--GGUGGGGGAAAU--AUGAGGAAAAACAAG ((((.(((((((((............)))))).......---------.....((...-.........))....--))).))))....--............... ( -13.80) >DroYak_CAF1 6567 93 + 1 CUUCGACCAAGCGCUGUAAUAAAUUCGCGCUUUAAGA-----------AAAGAGUAAA-AAUAAAAUGACAAAAAGGGUGGGGGAAAUAUAUGAGGAAAAACAAG ((((.(((((((((............)))))).....-----------.....((...-.........))......))).))))..................... ( -13.60) >consensus CUUCGACCAAGCGCUGUAAUAAAUUCGCGCUUUAAAA_A_________AAAGAGUAAA_AAUAAAAUGACAAAAA_GGUGGGGGAAAU__AUGAGGAAAAACAAG ((((.(((((((((............)))))).....................((.............))......))).))))..................... (-13.52 = -13.52 + -0.00)


| Location | 10,532,485 – 10,532,585 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 86.71 |
| Mean single sequence MFE | -9.47 |
| Consensus MFE | -9.10 |
| Energy contribution | -9.10 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.67 |
| Structure conservation index | 0.96 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521043 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10532485 100 - 27905053 CUUGUUUUUCCUCAU--AUUUCCCCCACC-UCUUUGUCAUUUUAUUUUUUACUGUUUUUCUUUAUUU--UUUAAAGCGCGAAUUUAUUACAGCGCUUGGUCGAAG ...............--..(((..(((..-..............................((((...--..))))((((............)))).)))..))). ( -9.60) >DroEre_CAF1 7454 91 - 1 CUUGUUUUUCCUCAU--AUUUCCCCCACC--UUUUGUCAUUUUAUU-UUUACUCUUU---------UUUUUUAAAGCGCGAAUUUAUUACAGCGCUUGGUCGAAG ...............--..(((..((..(--....)..........-..........---------.......((((((............))))))))..))). ( -9.40) >DroYak_CAF1 6567 93 - 1 CUUGUUUUUCCUCAUAUAUUUCCCCCACCCUUUUUGUCAUUUUAUU-UUUACUCUUU-----------UCUUAAAGCGCGAAUUUAUUACAGCGCUUGGUCGAAG ...................(((..((...(.....)..........-..........-----------.....((((((............))))))))..))). ( -9.40) >consensus CUUGUUUUUCCUCAU__AUUUCCCCCACC_UUUUUGUCAUUUUAUU_UUUACUCUUU_________U_UUUUAAAGCGCGAAUUUAUUACAGCGCUUGGUCGAAG ...................(((..((((.......))....................................((((((............))))))))..))). ( -9.10 = -9.10 + -0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:15 2006