
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 10,529,180 – 10,529,274 |
| Length | 94 |
| Max. P | 0.995817 |

| Location | 10,529,180 – 10,529,274 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 90.57 |
| Mean single sequence MFE | -24.55 |
| Consensus MFE | -22.21 |
| Energy contribution | -22.21 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.98 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.62 |
| SVM RNA-class probability | 0.995817 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
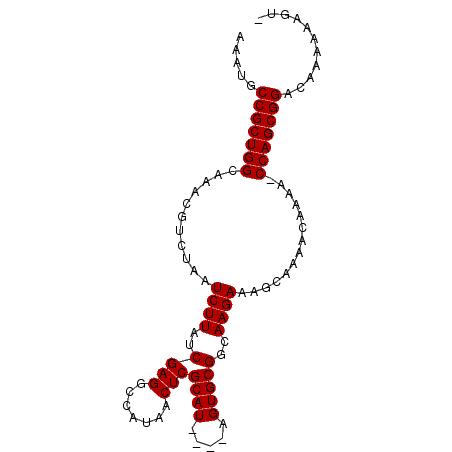
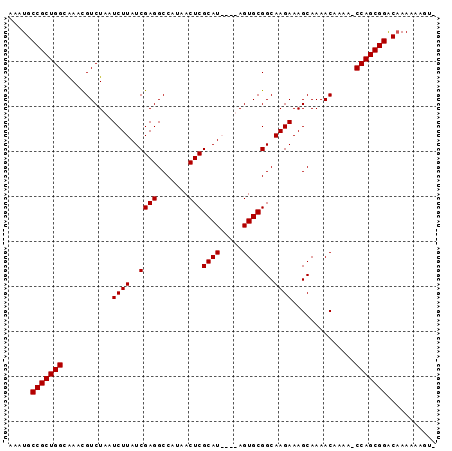
>3R_DroMel_CAF1 10529180 94 + 27905053 AAAUGCCGCUGGCAAACGUCUAAUCUUAUCGAGGCCAUAACUCGCAUACAUAGUGCGGCAAGAAAGCAAAACAAAA-CCAGCGGACAAAAGUAUA ...(((((((((....((((((....(((((((.......)))).)))..))).)))((......)).........-))))))).))........ ( -24.20) >DroSec_CAF1 4499 94 + 1 AAAUGCCGCUGGCAAACGUCUAAUCUUAUCGAGGCCAUAACUCGCAUACAUAGUGCGGCAAGAAAGCAAAACAAAA-CCAGCGGACUAAAAUGUA .....(((((((....((((((....(((((((.......)))).)))..))).)))((......)).........-)))))))........... ( -23.20) >DroEre_CAF1 4210 89 + 1 AAAUGCCGCUGGCGAACGUCUAAUCUUAUCGAGGCCAUAACUCGCAU----AGUGCGGCAAGAAAGCAAAACAAAA-CCAGCGGGCAAACAAGU- ...(((((((((..............(((((((.......)))).))----)((...((......))...))....-))))).)))).......- ( -23.20) >DroYak_CAF1 3208 90 + 1 AAAUGCCGCUGGCAAAUGUCUAAUCUUAUCGAGGCCAUAACUCGCAU----AGUGCGGCAAGAAAGCAAAACAAAAGCCAGCGGACAAAUAAGC- ...((((((((((.............(((((((.......)))).))----)((...((......))...))....)))))))).)).......- ( -27.60) >consensus AAAUGCCGCUGGCAAACGUCUAAUCUUAUCGAGGCCAUAACUCGCAU____AGUGCGGCAAGAAAGCAAAACAAAA_CCAGCGGACAAAAAAGU_ .....(((((((...........((((..((((.......)))((((.....)))))..))))..............)))))))........... (-22.21 = -22.21 + 0.00)

| Location | 10,529,180 – 10,529,274 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 90.57 |
| Mean single sequence MFE | -27.30 |
| Consensus MFE | -23.81 |
| Energy contribution | -23.62 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.78 |
| SVM RNA-class probability | 0.976737 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
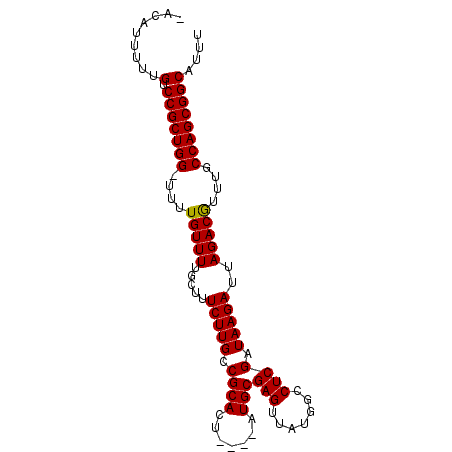
>3R_DroMel_CAF1 10529180 94 - 27905053 UAUACUUUUGUCCGCUGG-UUUUGUUUUGCUUUCUUGCCGCACUAUGUAUGCGAGUUAUGGCCUCGAUAAGAUUAGACGUUUGCCAGCGGCAUUU ........((.(((((((-(..(((((..(((((..((((.(((.((....)))))..))))...)).)))...)))))...))))))))))... ( -27.20) >DroSec_CAF1 4499 94 - 1 UACAUUUUAGUCCGCUGG-UUUUGUUUUGCUUUCUUGCCGCACUAUGUAUGCGAGUUAUGGCCUCGAUAAGAUUAGACGUUUGCCAGCGGCAUUU .........(.(((((((-(..(((((..(((((..((((.(((.((....)))))..))))...)).)))...)))))...))))))))).... ( -26.70) >DroEre_CAF1 4210 89 - 1 -ACUUGUUUGCCCGCUGG-UUUUGUUUUGCUUUCUUGCCGCACU----AUGCGAGUUAUGGCCUCGAUAAGAUUAGACGUUCGCCAGCGGCAUUU -.......((((.(((((-(..(((((..(((((..((((.(((----.....)))..))))...)).)))...)))))...))))))))))... ( -26.00) >DroYak_CAF1 3208 90 - 1 -GCUUAUUUGUCCGCUGGCUUUUGUUUUGCUUUCUUGCCGCACU----AUGCGAGUUAUGGCCUCGAUAAGAUUAGACAUUUGCCAGCGGCAUUU -.......((.((((((((...(((((..(((((..((((.(((----.....)))..))))...)).)))...)))))...))))))))))... ( -29.30) >consensus _ACAUUUUUGUCCGCUGG_UUUUGUUUUGCUUUCUUGCCGCACU____AUGCGAGUUAUGGCCUCGAUAAGAUUAGACGUUUGCCAGCGGCAUUU .........(.(((((((....(((((.....(((((.((((.......)))(((.......)))).)))))..)))))....)))))))).... (-23.81 = -23.62 + -0.19)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:05 2006