
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 10,522,160 – 10,522,319 |
| Length | 159 |
| Max. P | 0.636496 |

| Location | 10,522,160 – 10,522,266 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.88 |
| Mean single sequence MFE | -31.68 |
| Consensus MFE | -17.27 |
| Energy contribution | -17.58 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.636496 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10522160 106 - 27905053 ACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCAGUCGGAAGGAGAAAAGUCAAAAGGUGAGUUCCCUGAAAAGUCGCU-----------UAAGUCCGGGGGAAAUUCAAAUGC ...((((....(((((..((((((((((.((((.---((.....)).))))....)))...(((.(....)))).....))))))-----------)..).)))).)))).......... ( -30.80) >DroSec_CAF1 172402 108 - 1 ACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCAUCAUCAGUCGGAAGGAGAAAAGCCAAAAGGUGAGUUCCCGGAAAAGUCGCU-----------UAAGUU-GCGGGAAAAUCAAAUGA ...(((((((((......(((((((....((((....((.....)).((.(((..(((....)))...)))))))))..))))))-----------)..)))-))))))........... ( -34.50) >DroSim_CAF1 68983 105 - 1 ACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCAGUCGGAAGGAGAAAAGCCAAAAGGUGAGUUCCCGGAAAAGUCGCU-----------UAAGUU-GGGGGAAAAUCAAAUGA ...((((....(((((..(((((((....((((.---((.....)).((.(((..(((....)))...)))))))))..))))))-----------)..)))-)).)))).......... ( -32.60) >DroEre_CAF1 68443 103 - 1 ACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCAGUCGGAAGGAGAAAAGCCAAAAGGUGAGUUCCCGGAAAAGUAGCA-----------UAAGUC---GGAAGGUUUAAAUCC .....((((.((((((....))..)))).((((.---((.....)).((.(((..(((....)))...)))))))))..))))..-----------(((..(---....)..)))..... ( -30.60) >DroYak_CAF1 69376 94 - 1 ACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCAGUCGGAAGGAGAAAAGCCAAAAGGUGAGUUCCCGGAAAAGUU-----------------------GGGAAAAUCAAAUCC ...((((.(.((((((....))..)))).((((.---......))))).))))..(((....)))((.(((((((.....))-----------------------)))))..))...... ( -27.80) >DroAna_CAF1 57067 100 - 1 --------CUGCCAGCGGAAGCUGCCGCGG---------AGGGCGGCGGGGGAAAGCCAAGAGGUGAGUCGCCGGAAAAUGGGAAAACUCCCAAUAAAAGUC---UGAAAGUGCAUGUGC --------((((((((....))))..))))---------.((((..(((.((...(((....)))...)).))).....(((((....)))))......)))---).............. ( -33.80) >consensus ACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA___UCAGUCGGAAGGAGAAAAGCCAAAAGGUGAGUUCCCGGAAAAGUCGCU___________UAAGUC___GGGAAAAUCAAAUGC ..........((((((....))..)))).((((..........))))((.(((..(((....)))...)))))............................................... (-17.27 = -17.58 + 0.31)


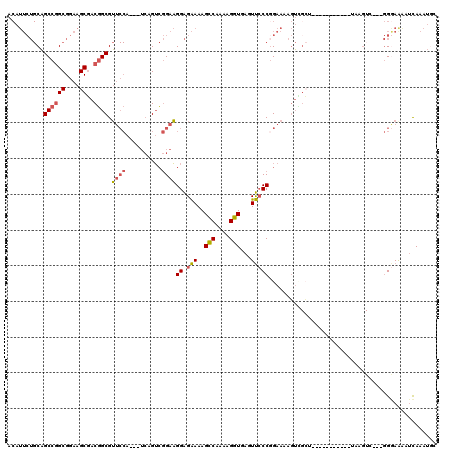
| Location | 10,522,229 – 10,522,319 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 82.56 |
| Mean single sequence MFE | -29.77 |
| Consensus MFE | -19.31 |
| Energy contribution | -20.32 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.65 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.512543 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10522229 90 + 27905053 UGA---UGGAACGCCGUCGCUUCCGCCGGCUGCAGAAUGUGUUGUUGCAAGCAUGUGUUGAGCAGCAGCAGCGGUAGCAAAAACAAAUGCAAG .((---(((....)))))(((.((((..(((((...(((((((......))))))).....)))))....)))).)))............... ( -31.30) >DroPse_CAF1 66985 78 + 1 U---------CAGCUGUGGCCUCCGCAA-CCCCACAG-----UGCUGCCAACAUAUGUUGAGCGAGAGCAACAACAGUGCCAGCAGUUGCAGC .---------..(((((((.........-..))))))-----)(((((.(((...(((((.((....))..)))))((....)).)))))))) ( -25.60) >DroSec_CAF1 172470 93 + 1 UGAUGAUGGAACGCCGUCGCUUCCGCCGGCUGCAGAAUGUGUUGUUGCAAGCAUGUGUUGAGCAGCAGCAGCGGUAGCAAAAACAAAUGCAAG ....(((((....)))))(((.((((..(((((...(((((((......))))))).....)))))....)))).)))............... ( -31.30) >DroSim_CAF1 69051 90 + 1 UGA---UGGAACGCCGUCGCUUCCGCCGGCUGCAGAAUGUGUUGUUGCAAGCAUGUGUUGAGCAGCAGCAGCGGUAGCAAAAACAAAUGCAAG .((---(((....)))))(((.((((..(((((...(((((((......))))))).....)))))....)))).)))............... ( -31.30) >DroEre_CAF1 68509 90 + 1 UGA---UGGAACGCCGUCGCUUCCGCCGGCUGCAGAAUGUGUUGUUGCAAGCAUGUGUUGAGCAGAAGCAGCGGUAGCAAAAACAAAUGCAAG .((---(((....)))))(((.((((.(.((((...(((((((......))))))).....))))...).)))).)))............... ( -27.80) >DroYak_CAF1 69433 90 + 1 UGA---UGGAACGCCGUCGCUUCCGCCGGCUGCAGAAUGUGUUGUUGCAAGCAUGUGUUGAGCAGCAGCAGCGGUAGCAGAAACAAAUGCAAG .((---(((....)))))(((.((((..(((((...(((((((......))))))).....)))))....)))).)))............... ( -31.30) >consensus UGA___UGGAACGCCGUCGCUUCCGCCGGCUGCAGAAUGUGUUGUUGCAAGCAUGUGUUGAGCAGCAGCAGCGGUAGCAAAAACAAAUGCAAG ............((((.((....)).))))((((.....(((((((((.(((....)))..)))))))))((....)).........)))).. (-19.31 = -20.32 + 1.00)



| Location | 10,522,229 – 10,522,319 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 82.56 |
| Mean single sequence MFE | -25.95 |
| Consensus MFE | -21.53 |
| Energy contribution | -21.07 |
| Covariance contribution | -0.47 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.572253 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10522229 90 - 27905053 CUUGCAUUUGUUUUUGCUACCGCUGCUGCUGCUCAACACAUGCUUGCAACAACACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCA .........((..(((((.((((((..(((((............................))))))))))).)))))..))......---... ( -25.79) >DroPse_CAF1 66985 78 - 1 GCUGCAACUGCUGGCACUGUUGUUGCUCUCGCUCAACAUAUGUUGGCAGCA-----CUGUGGGG-UUGCGGAGGCCACAGCUG---------A .(..((.((((..(((.(((((..((....)).)))))..)))..))))..-----.))..)((-(((.((...)).))))).---------. ( -27.50) >DroSec_CAF1 172470 93 - 1 CUUGCAUUUGUUUUUGCUACCGCUGCUGCUGCUCAACACAUGCUUGCAACAACACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCAUCAUCA .........((..(((((.((((((..(((((............................))))))))))).)))))..))............ ( -25.79) >DroSim_CAF1 69051 90 - 1 CUUGCAUUUGUUUUUGCUACCGCUGCUGCUGCUCAACACAUGCUUGCAACAACACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCA .........((..(((((.((((((..(((((............................))))))))))).)))))..))......---... ( -25.79) >DroEre_CAF1 68509 90 - 1 CUUGCAUUUGUUUUUGCUACCGCUGCUUCUGCUCAACACAUGCUUGCAACAACACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCA ...(((........)))..(((.((((((((((........(((.(((...........)))))).)))))))))).))).......---... ( -24.00) >DroYak_CAF1 69433 90 - 1 CUUGCAUUUGUUUCUGCUACCGCUGCUGCUGCUCAACACAUGCUUGCAACAACACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA---UCA ...((..(((((((((((...((((((((.((.........))..)))............))))).)))))))))))..))......---... ( -26.80) >consensus CUUGCAUUUGUUUUUGCUACCGCUGCUGCUGCUCAACACAUGCUUGCAACAACACAUUCUGCAGCCGGCGGAAGCGACGGCGUUCCA___UCA .........(((.(((((.((((((((((...........((....))............)))).)))))).))))).)))............ (-21.53 = -21.07 + -0.47)


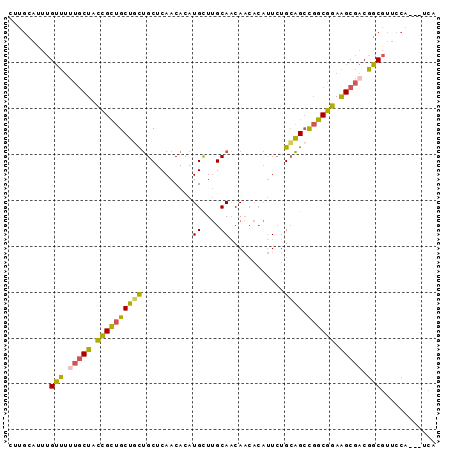
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:12:00 2006