
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 10,217,078 – 10,217,194 |
| Length | 116 |
| Max. P | 0.710164 |

| Location | 10,217,078 – 10,217,194 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 87.50 |
| Mean single sequence MFE | -38.20 |
| Consensus MFE | -27.76 |
| Energy contribution | -27.82 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553173 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10217078 116 + 27905053 GCUUUUGCAGACUCGAAUACAGUGUAGUGGCGCCACCUAUGCAGGCAUCACCGGCAAGCGCCACCUACACGGGCACUAGCAACUAGACUUGGCGCCAUCUACAACAACUGAACCGG ((....))...........((((((((((((((((.((((((((((......(((....))).((.....)))).)).)))..)))...)))))))).))))....))))...... ( -37.00) >DroSec_CAF1 75157 116 + 1 GCUUUUGCAGACUCGAAUACAGUGUAGUGGCGCCACCUAUGCAGGCAUCGCCGGCAAACGCCACCUACACGGGCAUUAGCAAUUAGAGUUGGCGCCAUCUACAACAACUGAACCGG ((....))...........(((((((((((((((((((((((.(((...)))(((....))).(((....))).....)))..))).).)))))))).))))....))))...... ( -37.80) >DroSim_CAF1 74977 116 + 1 GAUUUUGCACACUCAAAUACAAUGUAGUGGCGCCACCUAUGGAGCCAUAGCCGGCAUACGCCACCUACACGGGCCUUAGCAAUUUGAGUUGGCGCCAUCUACAACAACUGAACCGG (((..(((..((((((((.(..((((((((((((..(((((....)))))..)))....)))).)))))..)((....)).))))))))..)))..)))................. ( -34.50) >DroEre_CAF1 76931 115 + 1 GCUCUUGCAGGCUCAAAUACCAUGUAGUGGCGCCACCUGUGCAGGCAUCGCCAGCAGACGCCACCUGCACGGGCA-GAGCAAGUGGAUUGGGCGCCAUCUACAACAGCUGAAUCGG ..((..((.((........)).((((((((((((.((((((((((...((........))...))))))))))..-...(((.....)))))))))).)))))...)).))..... ( -43.50) >consensus GCUUUUGCAGACUCAAAUACAAUGUAGUGGCGCCACCUAUGCAGGCAUCGCCGGCAAACGCCACCUACACGGGCAUUAGCAAUUAGAGUUGGCGCCAUCUACAACAACUGAACCGG ......................((((((((((((.((.(((....)))....(((....)))........))((....))..........))))))).)))))............. (-27.76 = -27.82 + 0.06)



| Location | 10,217,078 – 10,217,194 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 87.50 |
| Mean single sequence MFE | -44.42 |
| Consensus MFE | -36.90 |
| Energy contribution | -36.90 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.710164 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10217078 116 - 27905053 CCGGUUCAGUUGUUGUAGAUGGCGCCAAGUCUAGUUGCUAGUGCCCGUGUAGGUGGCGCUUGCCGGUGAUGCCUGCAUAGGUGGCGCCACUACACUGUAUUCGAGUCUGCAAAAGC .(((..((((....((((.(((((((....((((...)))).(((..(((((((..((((....))))..)))))))..))))))))))))))))))...)))............. ( -45.20) >DroSec_CAF1 75157 116 - 1 CCGGUUCAGUUGUUGUAGAUGGCGCCAACUCUAAUUGCUAAUGCCCGUGUAGGUGGCGUUUGCCGGCGAUGCCUGCAUAGGUGGCGCCACUACACUGUAUUCGAGUCUGCAAAAGC .(((..((((....((((.((((((((.........((....))((((((((((..(((......)))..)))))))).))))))))))))))))))...)))............. ( -42.30) >DroSim_CAF1 74977 116 - 1 CCGGUUCAGUUGUUGUAGAUGGCGCCAACUCAAAUUGCUAAGGCCCGUGUAGGUGGCGUAUGCCGGCUAUGGCUCCAUAGGUGGCGCCACUACAUUGUAUUUGAGUGUGCAAAAUC .............(((((.((((((((.((......((((.((((.(((((.(...).))))).)))).)))).....)).)))))))))))))((((((......)))))).... ( -41.60) >DroEre_CAF1 76931 115 - 1 CCGAUUCAGCUGUUGUAGAUGGCGCCCAAUCCACUUGCUC-UGCCCGUGCAGGUGGCGUCUGCUGGCGAUGCCUGCACAGGUGGCGCCACUACAUGGUAUUUGAGCCUGCAAGAGC ........(((.((((((..(((((((((.....)))...-.(((.((((((((..((((....))))..)))))))).)))))))))(((....)))........)))))).))) ( -48.60) >consensus CCGGUUCAGUUGUUGUAGAUGGCGCCAACUCUAAUUGCUAAUGCCCGUGUAGGUGGCGUUUGCCGGCGAUGCCUGCAUAGGUGGCGCCACUACACUGUAUUCGAGUCUGCAAAAGC .(((..(((....(((((.((((((((.........((....))(((((((((((.((((....)))).))))))))).))))))))))))))))))...)))............. (-36.90 = -36.90 + -0.00)

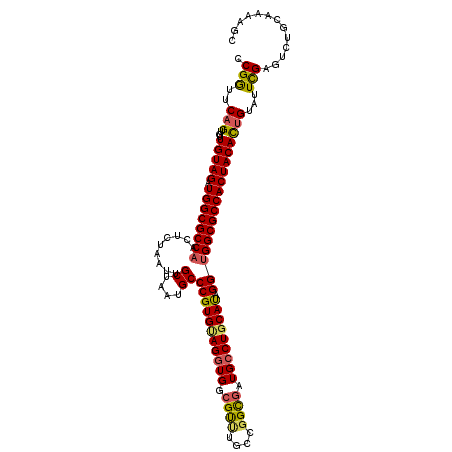

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:09:29 2006