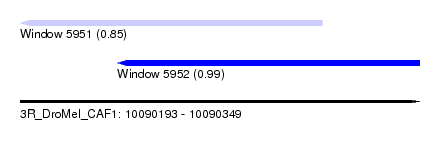

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 10,090,193 – 10,090,349 |
| Length | 156 |
| Max. P | 0.986233 |

| Location | 10,090,193 – 10,090,311 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 95.62 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -26.58 |
| Energy contribution | -26.82 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.849954 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10090193 118 - 27905053 ACCAAUUUUCAUUGCAAAAACUAAUUGCUGCCCACAAUGGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACUUGUAGUUAACGGUUAUAUCGAUCUAUGUUUGGUCAAUCG ..(((((((((((.((...........((((((.......))))))............)).)))))))))))...((..(((((....((((...))))..)))))..))........ ( -27.90) >DroSec_CAF1 148926 118 - 1 ACCAAUUUUCAUUGCAAAAAUUAAUUGCUGCCCACAAUAGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACUUGUAGUUAACGGUUAUAUCGAUCUAUGUUUGGUCAAUCG ..(((((((((((.((...........((((((.......))))))............)).)))))))))))...((..(((((....((((...))))..)))))..))........ ( -27.90) >DroEre_CAF1 161207 118 - 1 CCCAAUUUUCAUUGCAUAAACUAAAAGCUGCCCACAAUGGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACGUGUAGUUAACGGUUAUAUCGAUCUAUGUUUGGUCAAUCG ..(((((((((((.((...........((((((.......))))))............)).))))))))))).(((.(((.......)))))).....((((.......))))..... ( -30.70) >DroYak_CAF1 156150 118 - 1 UCCAAUUUUCAUUGCAAAAACUAAUAGCUGCCCACAAUAGAGGCAGAUAAAACUCACCUGGAAUGAGAAUUGAAACACUUGUAGUUAACGGUUAUAUCGAUCUAUGUUUGGUCAAUCG ..(((((((((((.((...........((((((......).)))))............)).)))))))))))...((..(((((....((((...))))..)))))..))........ ( -24.70) >consensus ACCAAUUUUCAUUGCAAAAACUAAUAGCUGCCCACAAUAGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACUUGUAGUUAACGGUUAUAUCGAUCUAUGUUUGGUCAAUCG ..(((((((((((.((...........((((((.......))))))............)).)))))))))))...((..(((((....((((...))))..)))))..))........ (-26.58 = -26.82 + 0.25)



| Location | 10,090,231 – 10,090,349 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.53 |
| Mean single sequence MFE | -28.75 |
| Consensus MFE | -25.30 |
| Energy contribution | -25.55 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.62 |
| Structure conservation index | 0.88 |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.986233 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 10090231 118 - 27905053 AAUCAAACAUGCUUGUA--UGUGCAAAGAUUAUAUUCUGUACCAAUUUUCAUUGCAAAAACUAAUUGCUGCCCACAAUGGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACUU ((((..(((((....))--))).....))))......(((..(((((((((((.((...........((((((.......))))))............)).)))))))))))..)))... ( -28.90) >DroSec_CAF1 148964 120 - 1 AGUCAAACUGGCUUGUAAUUGUGCAAAGAUUAUUUUCUGUACCAAUUUUCAUUGCAAAAAUUAAUUGCUGCCCACAAUAGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACUU ((((.....))))(((....(((((.(((....))).)))))(((((((((((.((...........((((((.......))))))............)).)))))))))))..)))... ( -30.70) >DroEre_CAF1 161245 120 - 1 AGUACAACAGGCUUGUAAUUGUGCAAAGAUUAUUUUCUGUCCCAAUUUUCAUUGCAUAAACUAAAAGCUGCCCACAAUGGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACGU .((((((...........))))))..(((......)))((..(((((((((((.((...........((((((.......))))))............)).)))))))))))..)).... ( -30.00) >DroYak_CAF1 156188 120 - 1 AGUCAAACAGGCCUGUAAUUGUGCAAAGAUUAUUUUCUGCUCCAAUUUUCAUUGCAAAAACUAAUAGCUGCCCACAAUAGAGGCAGAUAAAACUCACCUGGAAUGAGAAUUGAAACACUU .......((((...((((((.......))))))..))))...(((((((((((.((...........((((((......).)))))............)).)))))))))))........ ( -25.40) >consensus AGUCAAACAGGCUUGUAAUUGUGCAAAGAUUAUUUUCUGUACCAAUUUUCAUUGCAAAAACUAAUAGCUGCCCACAAUAGGGGCAGAAAAAACUCACCUGGAAUGAGAAUUGAAACACUU ....................(((...(((......)))....(((((((((((.((...........((((((.......))))))............)).)))))))))))...))).. (-25.30 = -25.55 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:08:12 2006