
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,842,824 – 9,842,949 |
| Length | 125 |
| Max. P | 0.989900 |

| Location | 9,842,824 – 9,842,922 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 86.45 |
| Mean single sequence MFE | -22.68 |
| Consensus MFE | -19.12 |
| Energy contribution | -19.62 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.74 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.989900 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9842824 98 - 27905053 CAUCCGUGCACAUUGGAAGGGCAUUGUGCUACUGCAAAUGCCCCAUUAGCUUUGUCUCUCUAUUUCCCCAUUCUAUUUGCCCAUUUCGAAGCUAC----AUA ..((((.......)))).(((((((.(((....))))))))))...((((((((................................)))))))).----... ( -23.45) >DroSec_CAF1 48869 102 - 1 GACCCGUGCACAUUGGAAGGGCAUUUUGCUACUGCAAAUGCCCCAUUAGCUUUGUCUCUCUAUUUCCCCAUUCUAUUUGCCCAUUUCGAAGCUACAUACAUA ...(((.......)))..((((((((.((....))))))))))...((((((((................................))))))))........ ( -21.85) >DroEre_CAF1 49880 96 - 1 CAUCCAUGCUCUUUGGAAGGGCAUUGAGCUACUGCACAGACCCCAUUAGCUUUGUGU--GUACUUCCCCAUUCUAUUUGCCCAUUUCAAAGCUAC----AAA ..........(((((((((((((..(((.(((.(((((((.(......).)))))))--))).)))...........))))).))))))))....----... ( -22.52) >DroYak_CAF1 50393 96 - 1 AACCCAUUCACAUUGGAAGGGCAUUGUGCUACUGCAAAUGCCCCAUUAGCUUUGUCU--CCACUUCCCCAAUCUAUUUGCCCAUUUCAAAGCUAC----AAA ...(((.......)))..(((((((.(((....))))))))))...((((((((...--...........................)))))))).----... ( -22.91) >consensus CACCCAUGCACAUUGGAAGGGCAUUGUGCUACUGCAAAUGCCCCAUUAGCUUUGUCU__CUACUUCCCCAUUCUAUUUGCCCAUUUCAAAGCUAC____AAA ...(((.......)))..(((((((.(((....))))))))))...((((((((................................))))))))........ (-19.12 = -19.62 + 0.50)



| Location | 9,842,852 – 9,842,949 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 83.10 |
| Mean single sequence MFE | -26.77 |
| Consensus MFE | -19.65 |
| Energy contribution | -19.84 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.73 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.529603 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9842852 97 + 27905053 UGGGGAAAUAGAGAGACAAAGCUAAUGGGGCAUUUGCAGUAGCACAAUGCCCUUCCAAUGUGCACGGAUGUAUAUUUUCUGAGAUGCCCUUCAAAAU .((((...(((((((...........(((((((((((....))).)))))))).....((((((....))))))))))))).....))))....... ( -27.00) >DroSec_CAF1 48901 97 + 1 UGGGGAAAUAGAGAGACAAAGCUAAUGGGGCAUUUGCAGUAGCAAAAUGCCCUUCCAAUGUGCACGGGUCUAUCUUUUCUGAGAUGCCCUCUAAAUU ...(((((.(((.((((...((....(((((((((((....)).)))))))))........))....)))).))))))))(((.....)))...... ( -27.60) >DroEre_CAF1 49908 95 + 1 UGGGGAAGUAC--ACACAAAGCUAAUGGGGUCUGUGCAGUAGCUCAAUGCCCUUCCAAAGAGCAUGGAUGUAUCUUUUGUUAGAUGCCCUAGAAAAU (((((...(((--(..((..(((...(((((.((.((....)).))..))))).......))).))..))))(((......)))..)))))...... ( -22.50) >DroYak_CAF1 50421 89 + 1 UGGGGAAGUGG--AGACAAAGCUAAUGGGGCAUUUGCAGUAGCACAAUGCCCUUCCAAUGUGAAUGGGUUUAUCUUUUCUGAGCUGCCCUA------ (((((.(((..--(((.((((.(((.(((((((((((....))).))))))))((((.......)))).))).)))))))..))).)))))------ ( -30.00) >consensus UGGGGAAAUAG__AGACAAAGCUAAUGGGGCAUUUGCAGUAGCACAAUGCCCUUCCAAUGUGCACGGAUGUAUCUUUUCUGAGAUGCCCUA_AAAAU .((((........(((.((((.((..(((((((((((....))).))))))))(((.........)))..)).)))))))......))))....... (-19.65 = -19.84 + 0.19)



| Location | 9,842,852 – 9,842,949 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 83.10 |
| Mean single sequence MFE | -24.07 |
| Consensus MFE | -17.28 |
| Energy contribution | -17.91 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.823090 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 9842852 97 - 27905053 AUUUUGAAGGGCAUCUCAGAAAAUAUACAUCCGUGCACAUUGGAAGGGCAUUGUGCUACUGCAAAUGCCCCAUUAGCUUUGUCUCUCUAUUUCCCCA .....((..((((................((((.......)))).(((((((.(((....)))))))))).........))))..)).......... ( -23.20) >DroSec_CAF1 48901 97 - 1 AAUUUAGAGGGCAUCUCAGAAAAGAUAGACCCGUGCACAUUGGAAGGGCAUUUUGCUACUGCAAAUGCCCCAUUAGCUUUGUCUCUCUAUUUCCCCA ........(((.......(((((((.(((((((.......)))..((((((((.((....))))))))))..........)))).))).))))))). ( -26.31) >DroEre_CAF1 49908 95 - 1 AUUUUCUAGGGCAUCUAACAAAAGAUACAUCCAUGCUCUUUGGAAGGGCAUUGAGCUACUGCACAGACCCCAUUAGCUUUGUGU--GUACUUCCCCA .......(((((((..................)))))))..(((((..(((.((((((.((.........)).))))))...))--)..)))))... ( -22.17) >DroYak_CAF1 50421 89 - 1 ------UAGGGCAGCUCAGAAAAGAUAAACCCAUUCACAUUGGAAGGGCAUUGUGCUACUGCAAAUGCCCCAUUAGCUUUGUCU--CCACUUCCCCA ------..(((.((...(((((((.(((..(((.......)))..(((((((.(((....))))))))))..))).)))).)))--...)).))).. ( -24.60) >consensus AUUUU_UAGGGCAUCUCAGAAAAGAUACACCCAUGCACAUUGGAAGGGCAUUGUGCUACUGCAAAUGCCCCAUUAGCUUUGUCU__CUACUUCCCCA ........(((......(((((((......(((.......)))..(((((((.(((....))))))))))......)))).))).........))). (-17.28 = -17.91 + 0.63)

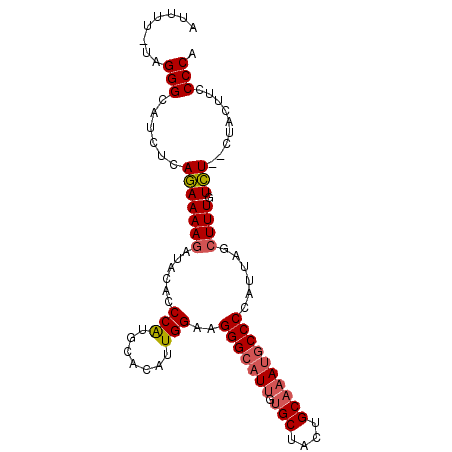

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:06:27 2006