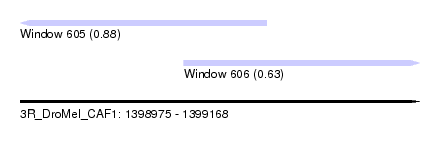
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 1,398,975 – 1,399,168 |
| Length | 193 |
| Max. P | 0.878901 |

| Location | 1,398,975 – 1,399,094 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.58 |
| Mean single sequence MFE | -43.12 |
| Consensus MFE | -38.01 |
| Energy contribution | -38.70 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.878901 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
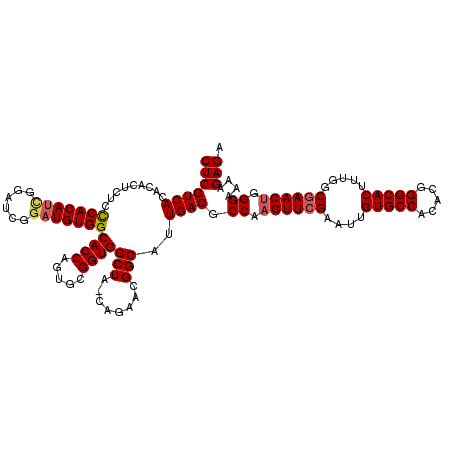
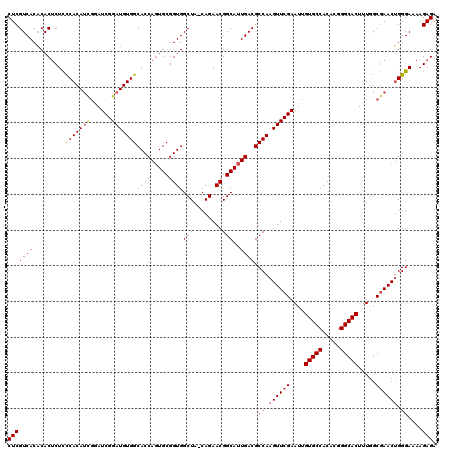
>3R_DroMel_CAF1 1398975 119 - 27905053 CUCGUCACACACUCUUCCACAUUGGAUCGGAUGUGGCACCAGUGCGGUGGCUA-CAGAACGGCAUUGACGCCAAGUUCGAAUUGUGCCACACGGGCACUUUGGCGAACUGGGAAAAGAGA (((((((((..(...(((.....)))..)..))))))..((((((.((.....-....)).))))))...((.((((((....(((((.....))))).....)))))).))....))). ( -43.40) >DroSec_CAF1 3804 119 - 1 CUCGUCACACACUCUCCCACAUCGAAUCGGAUGUGGCACCAGUGCGGUGGCUA-CAGAACGGCAUUGACGCCAAGUUCGAAUUGUGCCACACGGGCACUUUGGCGAACUGGGAAAAGAGA ...........((((((((((((......)))))(((..((((((.((.....-....)).))))))..))).((((((....(((((.....))))).....))))))))))...))). ( -44.30) >DroSim_CAF1 3739 119 - 1 CUCGUCACACACUCUCCCACAUCGGAUCGGAUGUGGCACCAGUGCGGUGGCUA-CAGAACGGCAUUGACGCCAAGUUCGAAUUGUGCCACACGGGCACUUUGGCGAACUGGGAAAAGAGA ...........((((((((((((......)))))(((..((((((.((.....-....)).))))))..))).((((((....(((((.....))))).....))))))))))...))). ( -44.30) >DroYak_CAF1 4043 117 - 1 CUCGUCACCCACACUCUCACAC---AUCGGAUGUGGCACCACUGCGGUGGCUCCCAGAACGGCAUUGACGCCAAGUUCGAAUUGUGCCACACGGGCACUUUAGCAAACGGGGAAAAGAGA (((((((((((((.(((.....---...)))))))(((....)))))))))((((.(((((((......)))..)))).....(((((.....)))))...........))))...))). ( -40.50) >consensus CUCGUCACACACUCUCCCACAUCGGAUCGGAUGUGGCACCAGUGCGGUGGCUA_CAGAACGGCAUUGACGCCAAGUUCGAAUUGUGCCACACGGGCACUUUGGCGAACUGGGAAAAGAGA (((((((.........(((((((......)))))))((((.....))))(((........)))..)))).((.((((((....(((((.....))))).....)))))).))....))). (-38.01 = -38.70 + 0.69)

| Location | 1,399,054 – 1,399,168 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 94.15 |
| Mean single sequence MFE | -39.00 |
| Consensus MFE | -34.27 |
| Energy contribution | -35.17 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.628342 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
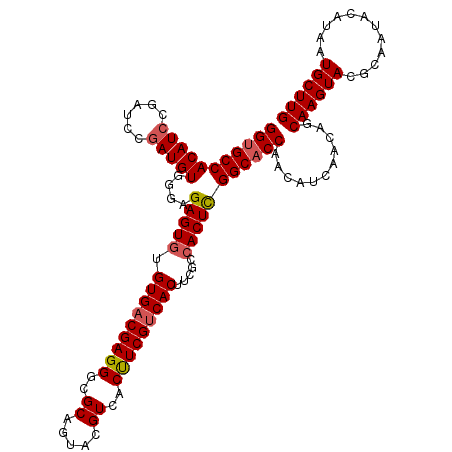
>3R_DroMel_CAF1 1399054 114 + 27905053 GGUGCCACAUCCGAUCCAAUGUGGAAGAGUGUGUGACGAGGGCGCAGUACGUCACUUCGCCACAUCGCAACUCGGCACCAACAUCAACAGCAAGUACGCAAUACAUAAUGCUUG ((((((........(((.....))).((((.(((((.(..((((.(((.....))).)))).).)))))))))))))))...........((((((............)))))) ( -37.00) >DroSec_CAF1 3883 114 + 1 GGUGCCACAUCCGAUUCGAUGUGGGAGAGUGUGUGACGAGGGCGCAGUACGUCACUUCGUCACUUCUCCACUUGGCCCCAACAUCAACAGCAAGUACGCAAUACAUAAUGCUUG ((.((((((((......))))).((((((...(((((((((..((.....))..)))))))))))))))....))).))...........((((((............)))))) ( -36.80) >DroSim_CAF1 3818 114 + 1 GGUGCCACAUCCGAUCCGAUGUGGGAGAGUGUGUGACGAGGGCGCAGUACGUCACCUCGUCACUUCGCCACUCGGCACCAACAUCAACAGCAAGUACGCAAUACAUAAUGCUUG (((((((((((......)))))....(((((.(((((((((..((.....))..))))))))).....)))))))))))...........((((((............)))))) ( -43.20) >consensus GGUGCCACAUCCGAUCCGAUGUGGGAGAGUGUGUGACGAGGGCGCAGUACGUCACUUCGUCACUUCGCCACUCGGCACCAACAUCAACAGCAAGUACGCAAUACAUAAUGCUUG (((((((((((......)))))....(((((.(((((((((..((.....))..))))))))).....)))))))))))...........((((((............)))))) (-34.27 = -35.17 + 0.89)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:41:26 2006