
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 9,430,314 – 9,430,409 |
| Length | 95 |
| Max. P | 0.953516 |

| Location | 9,430,314 – 9,430,409 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.30 |
| Mean single sequence MFE | -16.43 |
| Consensus MFE | -8.78 |
| Energy contribution | -9.08 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.53 |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.953516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
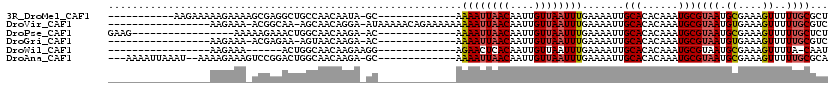
>3R_DroMel_CAF1 9430314 95 - 27905053 -----------AAGAAAAAGAAAAGCGAGGCUGCCAACAAUA-GC-------------AAAAUUAACAAUUGUUAAUUUGAAAAUUGCACACAAAUGCGUAAUGCGAAAGUUUUUGCGCU -----------.............((((.((((.......))-))-------------.((((((((....)))))))).....))))........((((((.((....))..)))))). ( -22.10) >DroVir_CAF1 105827 100 - 1 -----------------AAGAAA-ACGGCAA-AGCAACAGGA-AUAAAAACAGAAAAAAAAAUUAACAAUUGUUAAUUUGAAAAUUGCACACAAAUGCGUAAUGUGAAAGUUUUUGCGUC -----------------......-...((((-(((((..(..-.......)........((((((((....)))))))).....)))).((((.........))))......)))))... ( -13.60) >DroPse_CAF1 58091 89 - 1 GAAG-----------------AAAAAGAAACUGGCAACAAGA-AC-------------AAAAUUAACAAUUGUUAAUUUGAAAAUUGCACACAAAUGCGUAAUGCGAAAGUUUUUGCUCU ..((-----------------(...(((((((.((.......-..-------------.((((((((....))))))))....(((((.((....)).)))))))...)))))))..))) ( -15.80) >DroGri_CAF1 104397 87 - 1 -----------------AAGAAA-ACGAGAA-AGUAACAAGA-AC-------------AAAAUUAACAAUUGUUAAUUUGAAAAUUGCACACAAAUGCGUAAUGUGAAAGUUUUUGCGUC -----------------......-.......-.....(((((-((-------------.((((((((....))))))))....(((((.((....)).)))))......))))))).... ( -14.10) >DroWil_CAF1 63813 83 - 1 -----------------AAGAAA------ACUGGCAACAAGAAGG-------------AGAACUCACAAUUGUUAAUUUGAAAAUUGCACACAAAUGCGUAAUGCGAAAGUUUUA-CAAU -----------------...(((------(((...(((((....(-------------(....))....)))))..((((...(((((.((....)).))))).)))))))))).-.... ( -12.80) >DroAna_CAF1 56495 101 - 1 ---AAAAUUAAAU--AAAAGAAAGUCCGGACUGGCAACAAGA-GC-------------AAAAUUAACAAUUGUUAAUUUGAAAAUUGCACACAAAUGCGUAAUGCGAAAGUUUUUGCGCA ---..........--....(..(((....)))..)..(((..-..-------------.((((((((....)))))))).....)))........(((((((.((....))..))))))) ( -20.20) >consensus _________________AAGAAA_ACGAGACUGGCAACAAGA_AC_____________AAAAUUAACAAUUGUUAAUUUGAAAAUUGCACACAAAUGCGUAAUGCGAAAGUUUUUGCGCU ...........................................................((((((((....)))))))).......(((......)))((((.((....))..))))... ( -8.78 = -9.08 + 0.31)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:02:28 2006