
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 1,350,007 – 1,350,099 |
| Length | 92 |
| Max. P | 0.834947 |

| Location | 1,350,007 – 1,350,099 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 84.39 |
| Mean single sequence MFE | -18.70 |
| Consensus MFE | -14.54 |
| Energy contribution | -14.77 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.78 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
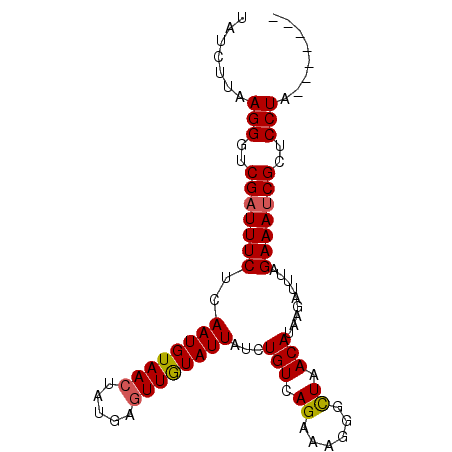
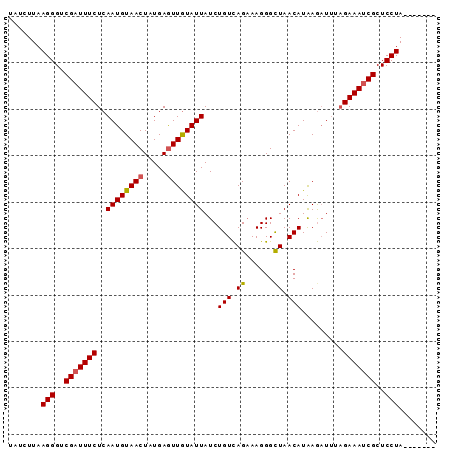
>3R_DroMel_CAF1 1350007 92 + 27905053 UAUCUUAAGGGACGUUUUCUCAAUGUAACUAUGAAUUAUAUUAUCUGUUAGUAAGGGCUAACAUGAACUAAUGAAAUCGCUCCUAAAUAUAG .......(((..((.((((..(((((((.......))))))).(((((((((....))))))).))......)))).))..)))........ ( -18.30) >DroSec_CAF1 14002 85 + 1 UAUCUUAAGGGUCGAUUUCUCAAUGUAACUCUGAGUUGUAUUAUCUGUCAGAAAGGGUUAACAUAAGAUUUAGAAAUCGCUCCUA------- .......(((..((((((((..((((((((((...(((.((.....)))))..)))))).)))).......))))))))..))).------- ( -20.40) >DroSim_CAF1 14025 85 + 1 UAUCUUAAGGGUCGAUUUCUCAAUGUAACUAUGAGUUGUAUUAUCUGUCAGAAAGGGCUAACAUAAGAUUUAGAAAUCGCUCCUA------- .......(((..((((((((.((((((((.....))))))))....(((......))).............))))))))..))).------- ( -17.40) >consensus UAUCUUAAGGGUCGAUUUCUCAAUGUAACUAUGAGUUGUAUUAUCUGUCAGAAAGGGCUAACAUAAGAUUUAGAAAUCGCUCCUA_______ .......(((..(((((((..((((((((.....))))))))...(((.((......)).))).........)))))))..)))........ (-14.54 = -14.77 + 0.23)

| Location | 1,350,007 – 1,350,099 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 84.39 |
| Mean single sequence MFE | -17.24 |
| Consensus MFE | -15.01 |
| Energy contribution | -15.23 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.834947 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
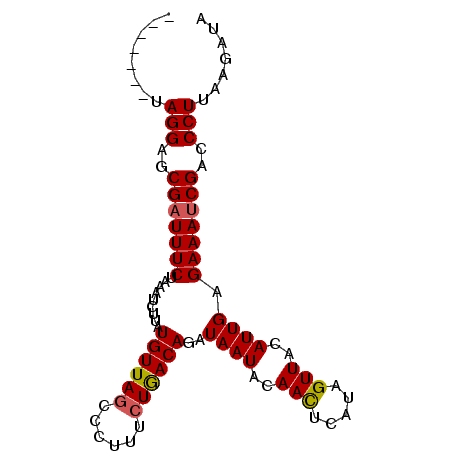
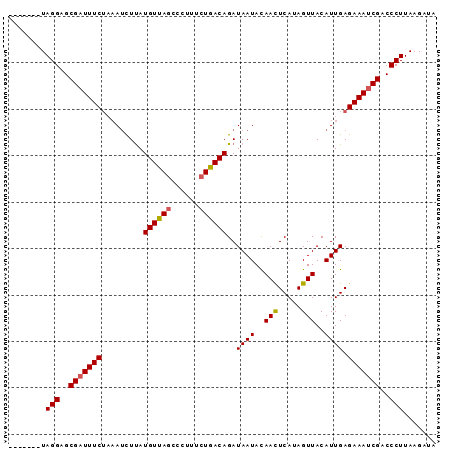
>3R_DroMel_CAF1 1350007 92 - 27905053 CUAUAUUUAGGAGCGAUUUCAUUAGUUCAUGUUAGCCCUUACUAACAGAUAAUAUAAUUCAUAGUUACAUUGAGAAAACGUCCCUUAAGAUA ...((((((((..((.((((......((.((((((......))))))))((((.((((.....)))).)))).)))).))..)))..))))) ( -16.30) >DroSec_CAF1 14002 85 - 1 -------UAGGAGCGAUUUCUAAAUCUUAUGUUAACCCUUUCUGACAGAUAAUACAACUCAGAGUUACAUUGAGAAAUCGACCCUUAAGAUA -------.(((..((((((((.......((((.(((....(((((.............))))))))))))..))))))))..)))....... ( -17.42) >DroSim_CAF1 14025 85 - 1 -------UAGGAGCGAUUUCUAAAUCUUAUGUUAGCCCUUUCUGACAGAUAAUACAACUCAUAGUUACAUUGAGAAAUCGACCCUUAAGAUA -------.(((..((((((((........((((((......))))))..((((..(((.....)))..))))))))))))..)))....... ( -18.00) >consensus _______UAGGAGCGAUUUCUAAAUCUUAUGUUAGCCCUUUCUGACAGAUAAUACAACUCAUAGUUACAUUGAGAAAUCGACCCUUAAGAUA ........(((..(((((((.........((((((......))))))..((((..(((.....)))..)))).)))))))..)))....... (-15.01 = -15.23 + 0.23)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:58 2006